Difference between revisions of "RNAi"
Schindelman (talk | contribs) |
Schindelman (talk | contribs) |
||
| Line 525: | Line 525: | ||
This web-based tool will automatically track papers that are positive based on our RNAi curation pipeline/checkout form and the OA; That is if a paper is on the checkout form and is curated, the data will be in the OA and marked as a True Positive. If the paper on the checkout list is a False Positive, the curator must note this on the paper editor (Flag False Positive) and the results will be forwarded to the SVM tracking form. | This web-based tool will automatically track papers that are positive based on our RNAi curation pipeline/checkout form and the OA; That is if a paper is on the checkout form and is curated, the data will be in the OA and marked as a True Positive. If the paper on the checkout list is a False Positive, the curator must note this on the paper editor (Flag False Positive) and the results will be forwarded to the SVM tracking form. | ||
http://tazendra.caltech.edu/~postgres/cgi-bin/paper_editor.cgi | http://tazendra.caltech.edu/~postgres/cgi-bin/paper_editor.cgi | ||
| − | The SVM lows that contain data are added to the checkout form (see above) and will be dealt with as any other paper on the list. The SVM lows that actually do not contain RNAi data must be entered manually onto the SVM tracking form. To do this go to | + | The SVM lows that do contain RNAi data are added to the checkout form by a curator (see above) and will be dealt with as any other paper on the list. The SVM lows that actually do not contain RNAi data must be entered manually onto the SVM tracking form as False Positives. To do this go to |
http://XXXXXXXXX.caltech.edu/~postgres/cgi-bin/svm_results.cgi | http://XXXXXXXXX.caltech.edu/~postgres/cgi-bin/svm_results.cgi | ||
Under "Enter Curator Results" | Under "Enter Curator Results" | ||
Revision as of 19:35, 20 August 2012
Contents
- 1 RNAi Curation Mission Summary
- 2 RNAi Data Model
- 3 RNAi Curation Standard Operating Procedure (SOP)
- 4 RNAi Paper Flagging and Processing of SVM "Low" Results and Tracking
- 5 RNAi OA
- 6 *.ACE Dumper Documentation
- 7 Parsing WS RNAi into OA
- 8 To sync to tazendra
RNAi Curation Mission Summary
The term "RNAi" stands for "RNA-interference" and refers to the targeted silencing of gene expression of a "target gene" via introduction of double stranded RNA (dsRNA) containing high degrees of sequence identity to the "target gene". In C. elegans, molecules of dsRNA may be introduced into the worm by a variety of different methods including direct micro-injection, soaking of worms in solution containing dsRNA, feeding worms bacteria that express dsRNA from a plasmid, and transgenic expression (within the worm) of dsRNA. For efficient and specific knockdown of gene expression, the dsRNA must have a minimum sequence identity with the target gene sequence. Any phenotype(s) resulting from the RNAi-mediated knockdown of a particular gene is thought to directly reflect the phenotype of a loss-of-function mutation in that gene, hence providing evidence as to the gene's biological function.
The goal of RNAi curation is to associate RNAi-mediated phenotypes with the target genes knocked down in RNAi experiments, as found in the literature pertaining to C. elegans and related species. There are some important things to consider when curating RNAi experiments. Some RNAi experiments are simple to curate as the strain is N2 (wild type genotype), only a single gene is targeted for RNAi, the phenotype is clearly stated, and the source and identity of the dsRNA is clearly stated by the authors. In many cases, however, the curation task is not so simple: there may be complex genotypes with multiple mutations in the strain receiving the dsRNA, multiple RNAi gene targets, complex genetic interactions, and/or missing descriptions from the authors as to controls, dsRNA delivery method, dsRNA identity, and/or phenotype directly resulting from the RNAi in question. Hopefully these issues will all be addressed below.
RNAi Data Model
This is the ?RNAi data model as of WormBase Release WS227:
//////////////////////////////////////////////////////////////////
//
// ?RNAi class
//
//////////////////////////////////////////////////////////////////
?RNAi Evidence #Evidence
History_name UNIQUE ?Text
Homol Homol_homol ?Homol_data XREF RNAi_homol ?Method Float Int UNIQUE Int Int UNIQUE Int #Homol_info
Sequence_info DNA_text Text UNIQUE Text //stores actual probe sequence for automated mapping
// 1st Text is DNA, 2nd is probe name
Sequence ?Sequence XREF RNAi //links to a real Sequence object used in the experiment
// such as yk clone; not UNIQUE anymore
PCR_product ?PCR_product XREF RNAi // links to a PCR_product object used in
// the experiment; not UNIQUE anymore
Uniquely_mapped //boolean; if present, signifies that ?RNAi object has a unique sequence
// which maps to a single place in the genome
Experiment Laboratory ?Laboratory
Author ?Author
Date UNIQUE DateType
Strain UNIQUE ?Strain
Genotype UNIQUE ?Text //used when no Strain object exists
Treatment UNIQUE ?Text
Life_stage UNIQUE ?Life_stage
Temperature UNIQUE Int
Delivered_by UNIQUE Bacterial_feeding //RL [010327]
Injection //RL [010327]
Soaking //RL [010327]
Transgene_expression //RL [010327]
Inhibits Predicted_gene ?CDS XREF RNAi_result #Evidence // "gene" parent (unreliable)
Gene ?Gene XREF RNAi_result #Evidence //RL [010327]
Transcript ?Transcript XREF RNAi_result #Evidence // [021126 krb]
Pseudogene ?Pseudogene XREF RNAi_result #Evidence // [030801 krb]
Supporting_data Movie ?Movie XREF RNAi // Lincoln, krb [010807]
DB_info Database ?Database ?Database_field ?Accession_number //to link out to Phenobank ar2 02-DEC-05
//removed UNIQUE as reqs multiple connections
Species UNIQUE ?Species
Gene_regulation ?Gene_regulation XREF RNAi // this tag is used when an RNAi experiment describes
// gene regulation ar2 29-MAR-06 for igor
Interaction ?Interaction
Reference UNIQUE ?Paper XREF RNAi //[070215 ar2] made reference unique so Paper sort of
// equates to a Study class for Will S
Phenotype ?Phenotype XREF RNAi #Phenotype_info
Phenotype_not_observed ?Phenotype XREF Not_in_RNAi #Phenotype_info //added by Wen to separate
// Not phenotype from real phenotypes
Expr_profile ?Expr_profile XREF RNAi_result // connection added during build [030106 krb]
Remark ?Text #Evidence
Method UNIQUE ?Method
Here is the WS231 ?Phenotype_info model:
////////////////////////////////////////////
//
// ?Phenotype_info Class
//
////////////////////////////////////////////
#Phenotype_info Paper_evidence ?Paper
Person_evidence ?Person
Curator_confirmed ?Person
Remark ?Text #Evidence // specific remarks about the phenotype
Quantity_description ?Text #Evidence //Remark to describe what quantity describes, below
Quantity UNIQUE Int UNIQUE Int #Evidence
Not #Evidence //This is being phased out but is needed for the next phase [06/08/10].
Anatomy_term ?Anatomy_term #Evidence
Penetrance Incomplete Text #Evidence
Low Text #Evidence
High Text #Evidence
Complete Text #Evidence
Range UNIQUE Int UNIQUE Int #Evidence // Range of penetrance
Recessive #Evidence
Semi_dominant #Evidence
Dominant #Evidence
Haplo_insufficient #Evidence
Caused_by ?Gene #Evidence
Caused_by_other ?Text #Evidence
Rescued_by_transgene ?Transgene
Loss_of_function UNIQUE Hypomorph #Evidence
Amorph #Evidence
Uncharacterised_loss_of_function #Evidence
Gain_of_function UNIQUE Dominant_negative #Evidence
Hypermorph #Evidence
Neomorph #Evidence
Uncharacterised_gain_of_function #Evidence
Other_allele_type UNIQUE Wild_type #Evidence
Isoallele #Evidence
Mixed #Evidence
Affected_by Molecule ?Molecule #Evidence // ?Molecule model Karen Yook
Temperature_sensitive Heat_sensitive Text #Evidence
Cold_sensitive Text #Evidence
Maternal UNIQUE Strictly_maternal #Evidence
With_maternal_effect #Evidence
Paternal #Evidence
Phenotype_assay Life_stage ?Life_stage #Evidence
Strain ?Strain #Evidence
Treatment ?Text #Evidence
Temperature ?Text #Evidence
Genotype ?Text #Evidence
Ease_of_scoring UNIQUE ES0_Impossible_to_score #Evidence
ES1_Very_hard_to_score #Evidence
ES2_Difficult_to_score #Evidence
ES3_Easy_to_score #Evidence
Here is the WS231 #Evidence hash model:
////////////////////////////////////////////////////////////////////////////////
// Evidence hash
////////////////////////////////////////////////////////////////////////////////
#Evidence Paper_evidence ?Paper // Data from a Paper
Published_as ?Text // .. track other names for the same data
Person_evidence ?Person // Data from a Person
Author_evidence ?Author UNIQUE Text // Data from an Author
Accession_evidence ?Database ?Accession_number // Data from a database (NDB/UNIPROT etc)
Protein_id_evidence ?Text // Reference a protein_ID
GO_term_evidence ?GO_term // Reference a GO_term
Expr_pattern_evidence ?Expr_pattern // Reference a Expression pattern
Microarray_results_evidence ?Microarray_results // Reference a Microarray result
RNAi_evidence ?RNAi // Reference a RNAi knockdown
Gene_regulation_evidence ?Gene_regulation // Reference a Gene_regulation interaction
CGC_data_submission // bless the data as comning from CGC
Curator_confirmed ?Person // bless the data manually
Inferred_automatically Text // bless the data via a script
Date_last_updated UNIQUE DateType // Stores last update timestamp
Feature_evidence ?Feature // Reference a Feature - eg for creation of isoform based on TEC-RED SL2
Laboratory_evidence ?Laboratory // Reference a Lab
From_analysis ?Analysis // Reference an analysis
Variation_evidence ?Variation // Explicitly record variation from which IMP manual GO annotations are made
Mass_spec_evidence ?Mass_spec_peptide
Sequence_evidence ?Sequence // for sequence data that hasn't been submitted to a public resource
RNAi Curation Standard Operating Procedure (SOP)
In order to ensure consistency of RNAi curation across curators and WormBase releases, a set of standard procedures for RNAi curation are outlined below. The descriptions of these procedures include use of the two main methods for generating *.ace files for release submission: (1) the web-based CGI form for one-at-a-time RNAi object generation and (2) the batch form submission method.
Minimum Requirements for an RNAi Object
Regardless of which curation method you (the curator) choose, there are a set of minimum requirements in order to generate a complete RNAi experiment object:
1) A reference (e.g. WBPaperID)
2) A curator name
3) The sequence of the dsRNA used in the experiment to knockdown gene expression
4) A dsRNA delivery method (e.g. Injection) [Note: some authors omit this information. This can be curated arbitrarily and removed from the *.ace file once it has been generated]
5) A phenotype (observed or not_observed)
If any one of these five basic pieces of information are missing, the scripts that generate the *.ace file will fail, returning an error.
Two Methods of Curation: Web CGI & Batch Form
There are two basic methods of RNAi curation: (1) a web-based CGI form and (2) a batch form involving a Perl script (living on the elbrus@caltech machine) to convert a spreadsheet (tab-delimited file) into an *.ace file.
(1) Web-based CGI
The web-based RNAi curation form can be found at:
http://elbrus.caltech.edu/cgi-bin/igor/rnaitools/rnai_curation
Web Form Page One
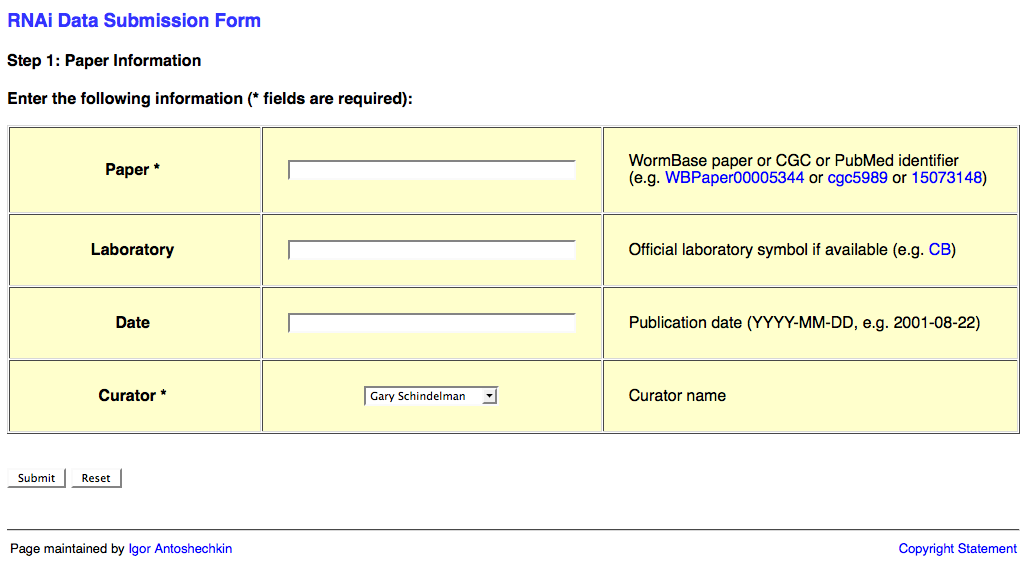
The first page of the web form requests information about the publication and looks like this:
This page requests information about the paper (e.g. WBPaperID), laboratory two-letter code (usually senior/corresponding author; for a list of all labs' two-letter codes look here: http://www.wormbase.org/db/misc/laboratory?name=*;class=laboratory), date of publication (accepted date), and a curator name. The laboratory and publication acceptance date are optional.
Once you've input the relevant information, click on the "Submit" button. Note that this may take a few moments to load. Once loaded you will be at the second page of the web form.
Web Form Page Two
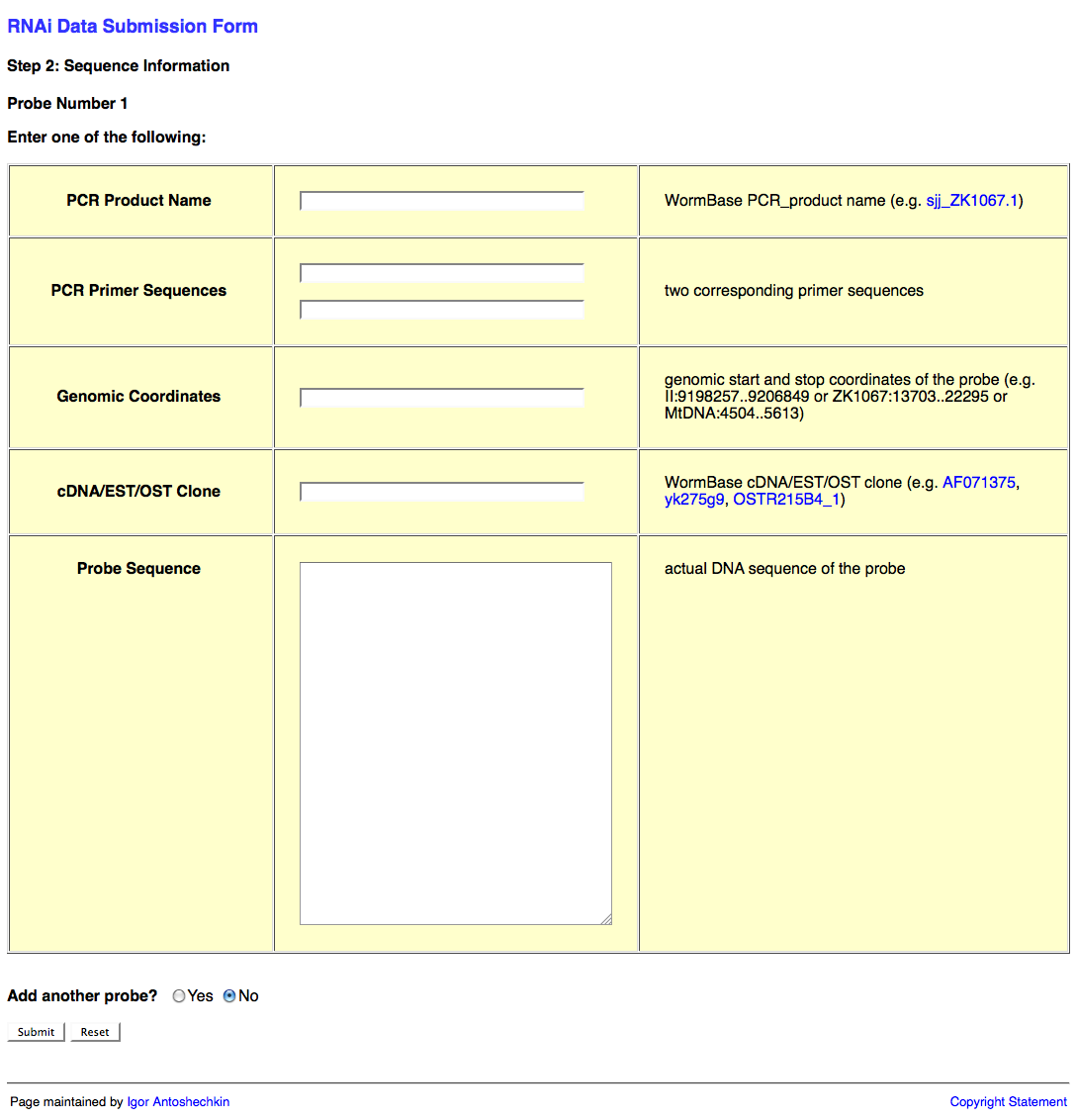
The second page of the web form looks like this:
This second page of the web form requests information about the RNAi probe(s) (i.e. the dsRNA sequence used to knockdown expression of the target gene). This information may be submitted in the form of a PCR product object (e.g. sjj_ZK617.1 or mv_CAA33463), PCR primers (it's usually good practice to check the primer sequences using e-PCR to confirm that they work properly), genomic coordinates, cDNA/EST/OST clone (note that many of these clones may be problematic; in such a case it is best to determine the sequence to the best of your ability and submit that instead), or actual sequence (e.g. "ACGT") of the probe.
An important point here is that RNAi target genes for an RNAi experiment are not submitted directly using the name of the target gene as indicated in the publication. The reason for this is that gene names can be continuously remapped to different regions of the genome over time, making persistence of the data impossible if we were to use only gene names to identify the target gene. By providing the actual sequence or clone used in the experiment, we can ensure that the appropriate gene is identified as the RNAi target gene regardless of which release of the database you are currently working in.
You may wish to submit more than one RNAi probe (e.g. when two or more genes are targeted by RNAi simultaneously). If this is the case, be sure to select "Yes" after the "Add another probe?" question at the bottom of the page, and then click on the "Submit" button.
Once all of your probes have been submitted, make sure that "No" is selected after the "Add another probe?" question, and click on the "Submit" button.
If all of the probes check out OK, you will be brought to the third page of the web form.
Web Form Page Three
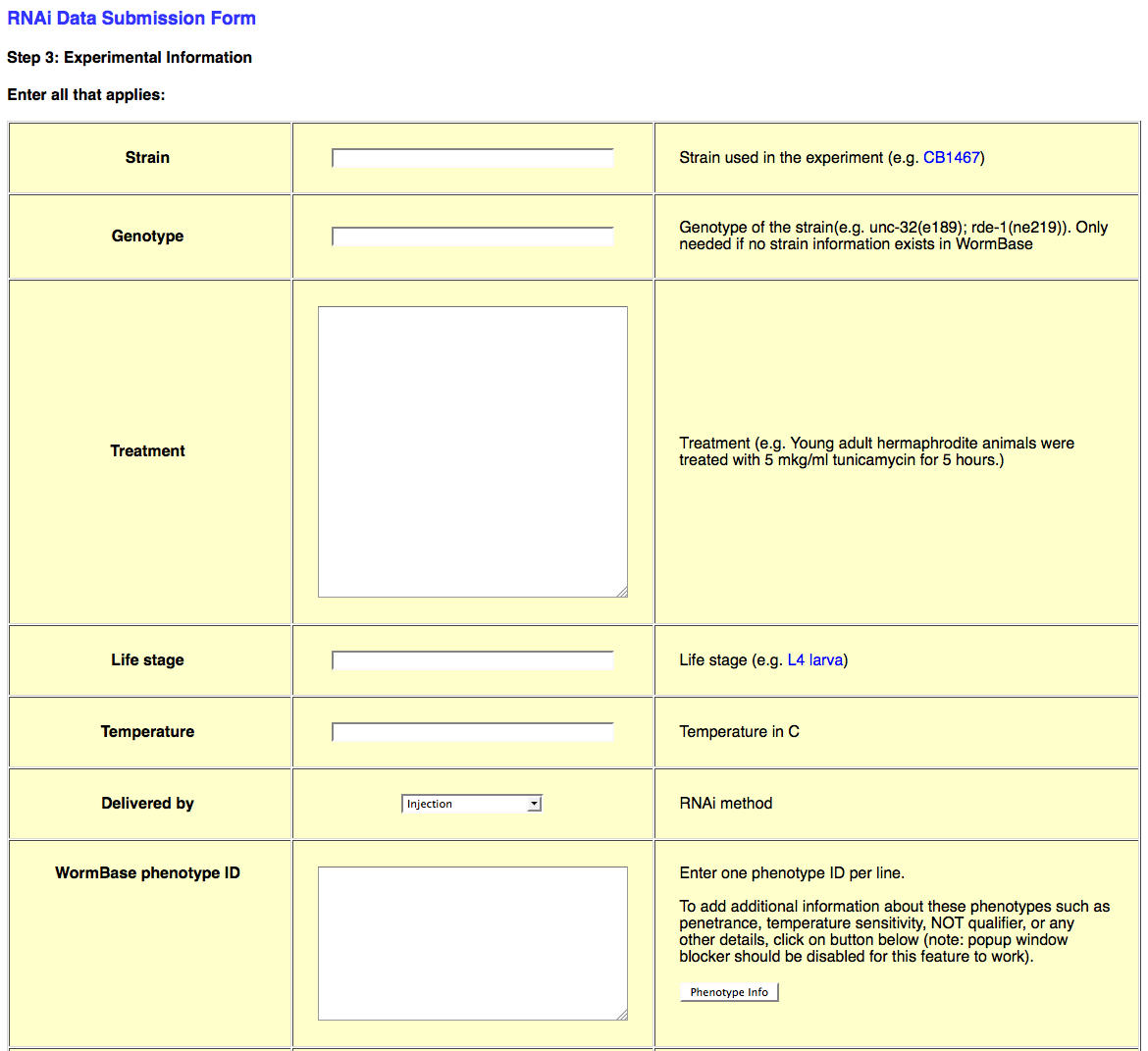
The third page of the web form looks like this:
This page requests all of the experimental details about the RNAi experiment in question. This information includes Strain/Genotype, treatment conditions, life stage, dsRNA delivery method (e.g. bacterial feeding), Phenotype observed (or not_observed), Gene Regulation objects, Genetic Interaction objects, species, and general remarks about the experiment.
Strain/Genotype
Whereas the "Genotype" information for a strain used in an RNAi experiment may be written in as free-text, the "Strain" field will only accept strains that are officially recognized by WormBase. Be sure to check the database for the existence of your strain name before submitting (otherwise you may receive an error upon submission). If your strain name does not currently exist in the database, you may write in the genotype in the "Genotype" field, or request the addition of the strain into the database.
Treatment/Temperature
The "Treatment" field is a free-text field in which the curator may specify any particular experimental conditions that apply to this RNAi experiment. This may include details about growth conditions, dsRNA delivery, or specimen manipulation. The "Temperature" field must be filled in with an integer value indicating the temperature in degrees Celsius (e.g. "25" for 25 degrees Celsius).
Life Stage
The "Life Stage" field must be filled in with an official life stage name, as stored in ACeDB/WormBase. For an official list of Life Stage names in WormBase, see:
or
Delivered by
The "Delivered by" field provides a drop-down menu of four dsRNA delivery methods: Injection, Bacterial feeding, Soaking, and Transgene expression.
WormBase phenotype ID
This field needs to have at least one phenotype in the form of a WormBase phenotype ID, for example: WBPhenotype:0000050 (for embryonic lethality). Without at least one phenotype, the *.ace-generating script will throw an error.
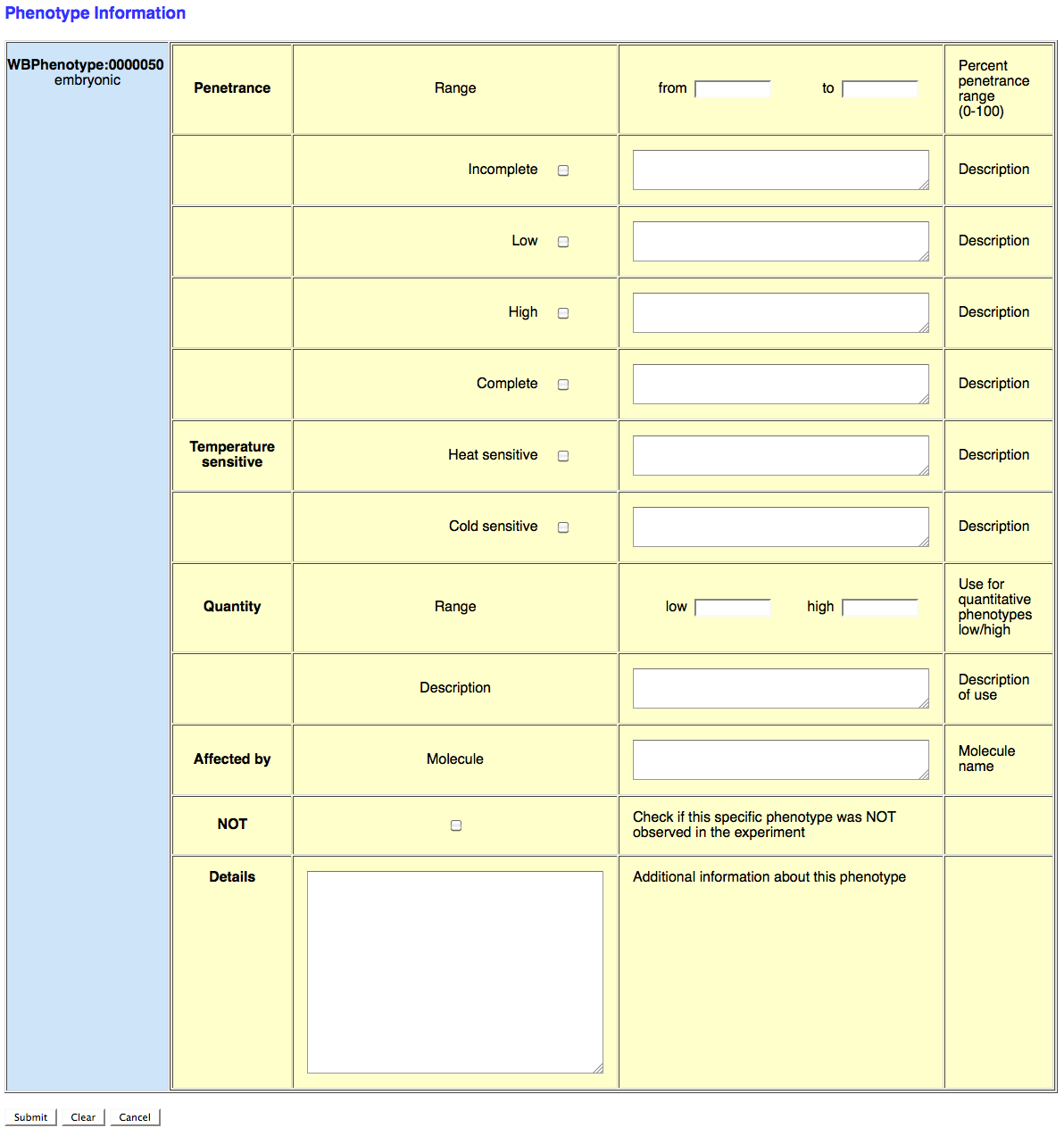
If you need to add other information pertaining to the phenotype, such as penetrance, quantification, or a remark, click on the "Phenotype Info" button to the right. This will open up a separate window that looks like this:
This form provides fields for the curator to enter information regarding the penetrance, temperature sensitivity, quantification, molecule interactions, phenotypes-not-observed, and a general, free-text remark field.
The Penetrance section records the range of penetrance of the phenotype in question (indicated to the left). The curator can provide a lower limit ("from") and/or an upper limit ("to"); if there is only one penetrance value indicated in the publication, just fill it in the "from" field. This value is an integer value representing percent penetrance. Below the "Range" fields, the curator can toggle the check boxes next to "Incomplete", "Low", "High", or "Complete" indicating the general quantity of penetrance. Each field also provides a free-text "Description" field to provide a description of the penetrance.
The Temperature sensitive fields allow the curator to indicate whether or not the phenotype is dependent on temperature and free-text "Description" field to provide details.
The Quantity fields record the lower and/or upper limits of the phenotype quantification as integer values on a scale that can be described in the "Description" free-text field. An example of a phenotype quantity might be "average brood size" or "body bends per minute".
The Affected by field captures molecules/compounds/drugs that affect the phenotype as indicated by this RNAi experiment. The molecule must be entered as the official WormBase molecule ID, usually the MeSH ID (e.g. D003609).
The NOT field provides a toggle check box for indicating if the phenotype in question was NOT OBSERVED in the experiment. This captures negative results of RNAi experiments.
Finally, the Details section provides a free-text field to provide descriptions of the phenotype beyond which could be captured in the other fields.
Additional phenotype information...
OBSOLETE; If you cannot find an appropriate phenotype for your RNAi experiment in the Worm Phenotype Ontology (WPO; worm_phenotype.obo can be downloaded from http://www.obofoundry.org/), please visit (http://tazendra.caltech.edu/~postgres/cgi-bin/new_objects.cgi) and enter the phenotype term you want to request by clicking the “Update Phenotype!” button and filling out the fields under “Suggest data through this CGI”.
New Gene Regulation
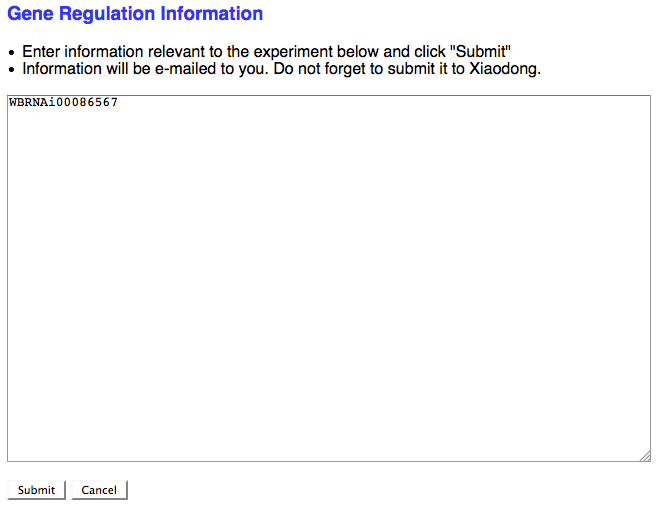
If the RNAi experimental result suggests that the RNAi target gene is somehow functionally involved in the regulation of a gene (at the transcriptional, translational, or post-translational level) and no Gene Regulation object exists for this relationship in the database, check the box here to request that a Gene Regulation object ID be assigned to this RNAi experiment. If the curator checks the box, a separate browser window will open with a simple text field to enter in the details of the Gene Regulation event:
As the note suggests, don't forget to send this info to Xiaodong so she can generate a new Gene Regulation Object.
Gene Regulation Object
If a Gene Regulation Object ID has already been assigned to this experiment from a different curation pipeline, enter the Gene Regulation Object ID here (e.g. WBPaper00006370_lin-3, cgc6303_lag-2).
New Interaction
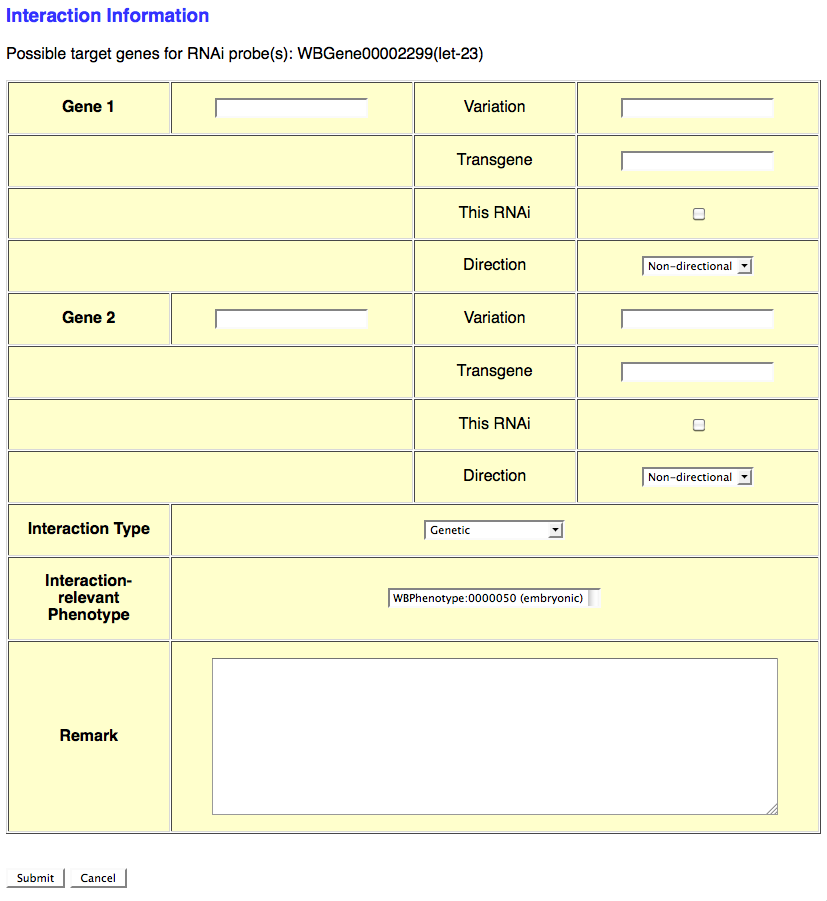
If the RNAi experiment indicates a genetic (or any other type) interaction and an Interaction Object ID has not yet been assigned to this interaction, select the number of genes involved in the interaction from the provided pull down menu. This will automatically open another browser window which will provide a form for entering details specific to that particular interaction (see image below):
This form provides the curator with fields to enter information for each interacting gene as well as general information about the interaction. The gene-specific information includes the gene's WBGeneID (e.g. WBGene00002299), "Variation" information (e.g. e1370, mgDf47) if this gene was perturbed by an allele or other variation, "Transgene" for a transgene harboring the gene (or variation of the gene) that contributes to the interaction, "This RNAi" toggle box to indicate that the perturbation of this gene was via RNAi (as opposed to an allele), and "Direction" drop-down menu to indicate if the gene is the "Effector" or "Effected" gene (for directional interactions) or "Non-directional" (for interactions in which directionality is irrelevant).
Below the gene-specific fields is a drop-down menu for indicating the "Interaction Type" (e.g. Genetic, Suppression, Enhancement), a "Interaction-relevant Phenotype" field to indicate the phenotype about which the interaction is based, and a free-text "Remark" field to describe any details of the interaction not sufficiently explained by the other fields.
Interaction Object
If an Interaction Object ID has already been assigned to this experiment from a different curation pipeline, enter the Interaction Object ID here (e.g. WBInteraction0001295).
Picture
OBSOLETE; The current RNAi data model no longer supports Picture objects
Species
Enter the species in which the RNAi experiment is taking place. Caenorhabditis elegans is set by default.
Remark
The "Remark" field is a free-text field to enter in any remaining relevant information that is pertinent to the RNAi experiment, but has not yet been captured by any other field of the curation form. This is often where standard remarks regarding the ambiguity of the dsRNA identity (see below) are entered. Information relevant to a genetic interaction, gene regulation event, or complex phenotype may also be put here.
If you are done curating this particular RNAi experiment and have more from the current paper, select "Yes" next to the "Add another experiment from this paper?" question at the bottom of the page and then click "Submit". If you are done curating RNAi experiments from this paper (or are done for the current session) and would like to generate an *.ace file, select "No" next to "Add another experiment from this paper?". Finally, indicate whether or not you would like to receive an e-mail of the *.ace file (as text in the body of the e-mail) next to "Would you like to receive a copy of the ace file via e-mail?". Once you are set, click the "Submit" button to generate the ace file (to be displayed in the web browser). To generate the *.ace file, you may simply copy and paste the text from the screen into a plain text file and save it with the ".ace" extension.
Post-processing your *.ace file
The .ace files from the CGI form need to be concatenated and processed using the following script (rnai_ace_modify.pl). Talk to a current RNAi curator for a copy of this script. This script converts the “NOT” tag to a “phenotype_not_observed” tag to be compliant with the current model and it splits the molecule data and interaction data into separate files: Hence, this script will take the .ace file from the CGI form and split it into 3 parts:
(1) FILENAME.ace.new -- for RNAi data
(2) FILENAME.ace.mol -- for molecule data
(3) FILENAME.ace.interaction - for interaction data
To run this script, type ./rnai_ace_modify.pl
It will ask you for the input FILENAME from the .ace file that you want to convert into these 3 files.
The following files get uploaded to CitaceMinus:
(1) FILENAME.ace.new -- for RNAi data
(2) FILENAME.ace.mol -- for molecule data
and the interaction file “FILENAME.ace.interaction” needs to be parsed into the interaction OA. (see the interaction Wiki page for instructions).
(2) Batch Form
The second basic method for generating RNAi *.ace files involves filling out all relevant RNAi curation information on a spreadsheet with each row (more or less; explanation below) as an RNAi experiment and each column as an RNAi object field. An Excel-based template file may be found here:
File:RNAi Curation Template.xls
Much of what applies to the RNAi Curation CGI Form described above applies similarly (or exactly) to the Batch Form. There are a few spreadsheet-specific changes that we will describe here.
First, there are a number of columns that are highlighted in yellow and have the comment "remove column before submission" in parentheses. These are temporary aids to the curator that must be removed before submitting the spreadsheet to the script, as they will cause an error.
Filling out the Batch Form
Shown below are screenshots from a fictional RNAi experiment entry in the batch form.
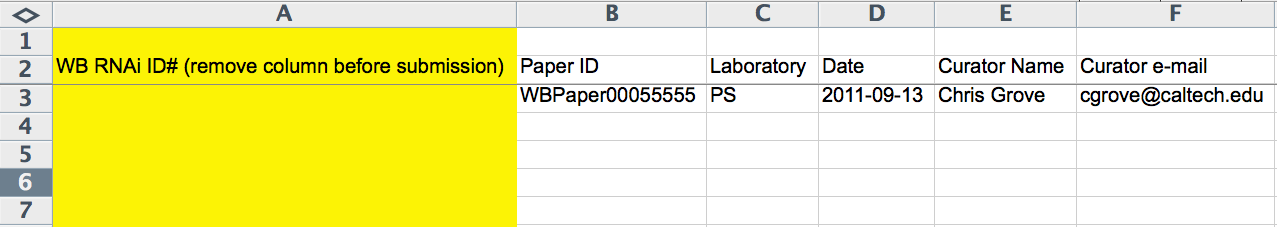
Paper Identification Information
The first 6 columns (A-F) of the spreadsheet record WBRNAi ID, WBPaper ID (WBPaper000#####), Laboratory (2-letter code), Publication Acceptance Date (YYYY-MM-DD), Curator Name, and Curator E-Mail. The WBRNAi ID column is only used once the *.ace file has already been made and is removed before submission to the *.ace-file-generating script. Whereas the Laboratory code and Publication Acceptance Date are optional, the WBPaper ID and the Curator Name are required.
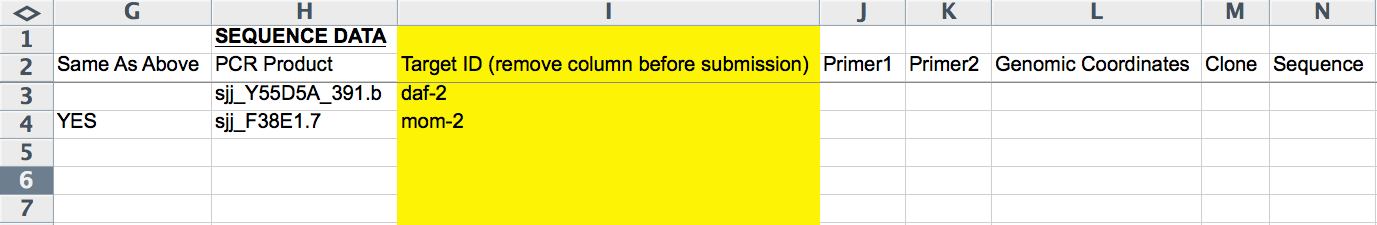
RNAi probe (dsRNA source) Information
The next 8 columns (G-N) record RNAi probe information (i.e. information about the dsRNA identity/sequence). The first column (G) is an exception. The "Same As Above" column either takes no entry or "YES" (all CAPS important), available from an Excel pull down menu; this column indicates whether the row/line of the form is to be included as part of the same RNAi experiment as the line(s) before it (the closest preceding line with paper information provided). This is necessary, for example, when submitting more than one RNAi probe, as each RNAi probe must go on its own line. It is important that information that is unique to an RNAi experiment (e.g. Strain/Genotype, Treatment, Temperature, etc.; indicated by RED text) is not also added to this line as it will overwrite any information provided in lines above it (belonging to the same RNAi experiment).
The "PCR Product", "Primer1", "Primer2", "Genomic Coordinates", "Clone", and "Sequence" columns record the dsRNA source as in the CGI Web Form (see above). The "Target ID" column (yellow, hence to be removed before submission) is free-text to indicate which gene the RNAi probe is targeting. This column is handy for keeping track of which RNAi experiments have already been curated, as there is no other easy way to see the gene's name in the form.
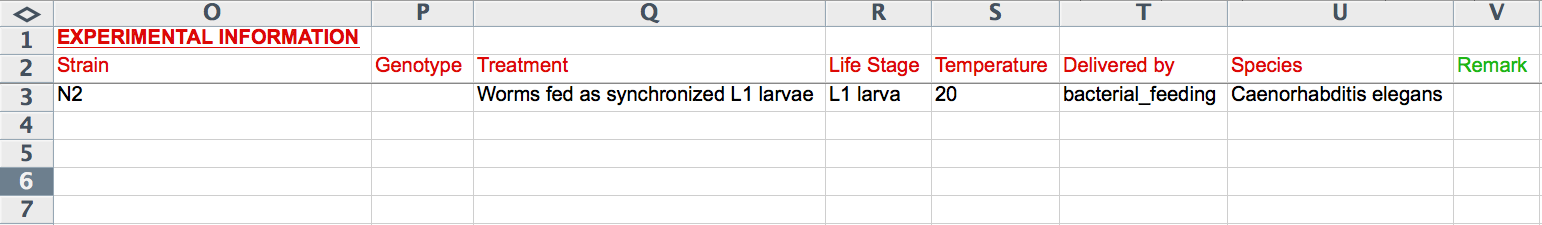
Experimental Information
The next 8 columns (O-V) record experimental details including "Strain", "Genotype", "Treatment", "Life Stage", "Temperature", "Delivered by", "Species", and "Remark". The "Life Stage" and "Delivered by" columns provide Excel pull down menus to select terms from a standard list (syntax is important). Note that these fields are the ones that are unique to an RNAi experiment; if two or more rows contain information for one of these columns, only the bottom-most row will be read. The formats for the remaining columns are as indicated in the CGI Web Form section above (i.e. "Genotype", "Treatment", and "Remark" are free-text, "Strain" must be a recognized WormBase strain name, "Temperature" must be an integer value for degrees Celsius, and "Species" must be a recognized Latin species name of a nematode species in WormBase).
Phenotype Information 1
Phenotype Information 2
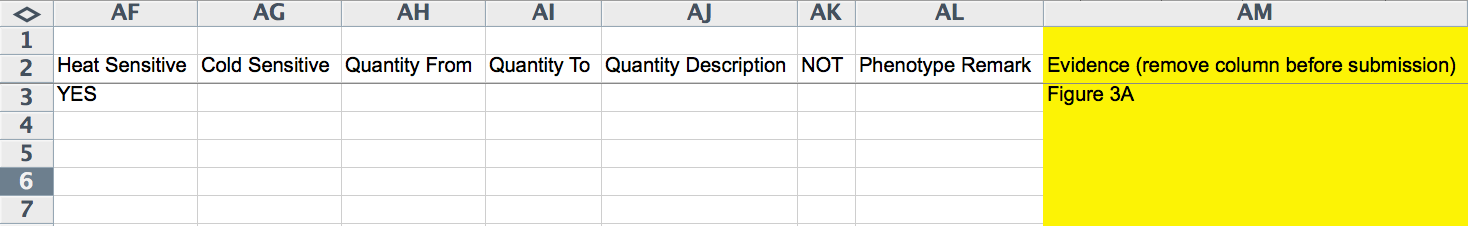
The next 17 columns (W-AM) record phenotype information. The "Gene Regulation ID" records a Gene Regulation Object ID, if one has already been generated for this experiment. The "Phenotype ID" must be given as the WormBase Phenotype ID as recorded in the Worm Phenotype Ontology (WPO; worm_phenotype.obo is available at http://www.obofoundry.org/), for example, WBPhenotype:0000081 for "L1 arrest". The yellow "Phenotype Name" column records the human-readable name of the phenotype for curatorial convenience.
The following columns (Z-AE) record the penetrance of the phenotype: "Penetrance From" records a lower limit of penetrance (integer of percentage), "Penetrance To" records an upper limit of penetrance (integer of percentage), and "Penetrancs(sic) Incomplete", "Penetrance Low", "Penetrance High", and "Penetrance Complete" are record ("YES" or empty; pull down menu) whether or not the penetrance quantification falls under any of these categories. "Heat Sensitive" and "Cold Sensitive" indicate ("YES" or empty; pull down menu) whether or not the phenotype is heat or cold sensitive, respectively. "Quantity From" and "Quantity To" capture the lower and/or upper limits of the phenotype quantification (integer values), if one is provided. "Quantity Description" is a free-text field for describing the nature of the phenotype quantitation. The "NOT" field would be filled with a "YES" if the phenotype was not observed (negative result). "Phenotype Remark" is a free-text field for describing any aspects of the phenotype not adequately captured by the other fields. The yellow "Evidence" field (optional and must be removed before submission) provides the curator with a field to capture the source of the evidence within the paper (e.g. Figures, Tables, Text, etc.).
Molecule Information
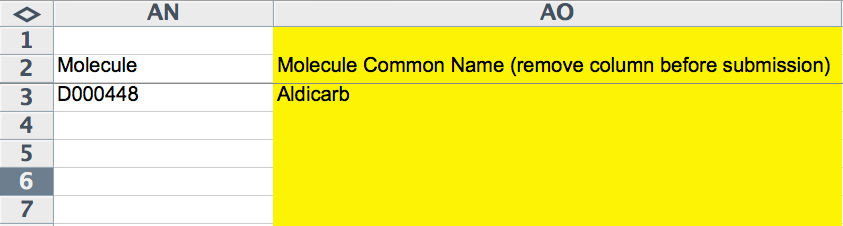
The next two columns (AN and AO) capture molecule information, if applicable. This would apply in circumstances in which a compound/drug was used in the experiment and potentially had biological activity or an affect on the phenotype being reported. The "Molecule" column must be filled with the WormBase recognized ID for a molecule, the MeSH ID by default. The yellow "Molecule Common Name" field should indicate (for curator convenience) the common, human-readable name of the compound.
Interaction Information 1
Interaction Information 2
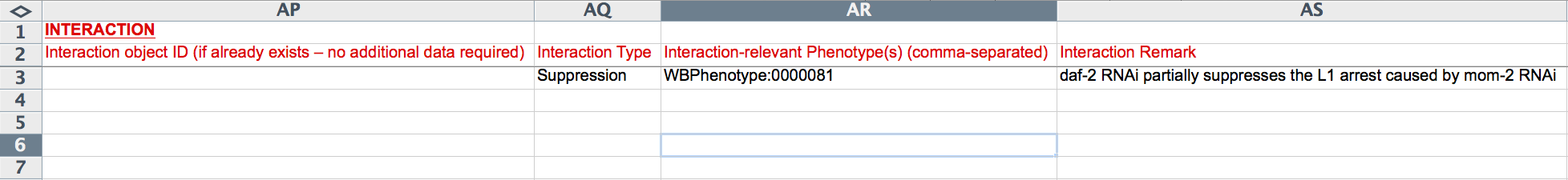
The last 10 columns (AP-AY) capture genetic interaction information. If a relevant Interaction Object ID has already been generated for this RNAi experiment, it may be entered in the "Interaction object ID" field, obviating the need to enter any further interaction information. If not, the following fields must be populated. The "Interaction Type" field provides a drop-down menu to select from a list of standard interaction type names (e.g. "Suppression"). The "Interaction-relevant Phenotype(s)" field records the phenotype ID of the phenotype about which the interaction is based, usually the same phenotype as was given under "Phenotype ID" (Column X). "Interaction Remark" is a free-text field describing the details of the interaction, for example, which gene suppressed which other gene with respect to what phenotype and under what conditions.
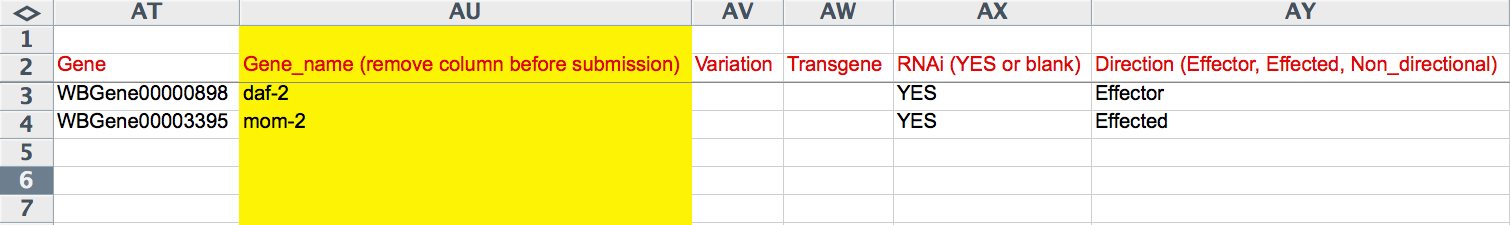
The "Gene" column reports the WormBase Gene ID (e.g. WBGene00000898) for the genes involved in the interaction. The yellow "Gene_name" column records the human-readable gene name of each gene involved in the interaction. The "Variation" records the allele/variation name of the gene involved in the genetic interaction, if applicable. This must be a recognized WormBase allele/variation name. If a transgene is involved in the interaction, the name of the transgene (official WormBase name) must be entered here. The "RNAi" column captures ("YES" or empty) whether or not the gene for that row was perturbed by RNAi in the interaction. "Direction" refers to the directionality of the interaction and the role that the gene (for that row) had in the interaction. This field provides a drop down list of flags: "Non_directional", "Effector", and "Effected". A suppressor/enhancer gene/mutation would be an "Effector" and the suppressed/enhanced gene/mutation would be the "Effected", whereas if the interaction is Non-directional (as in mutual enhancment, for example), each gene/mutation would be flagged as "Non_directional".
Submitting the Batch Form to Generate an *.ACE File
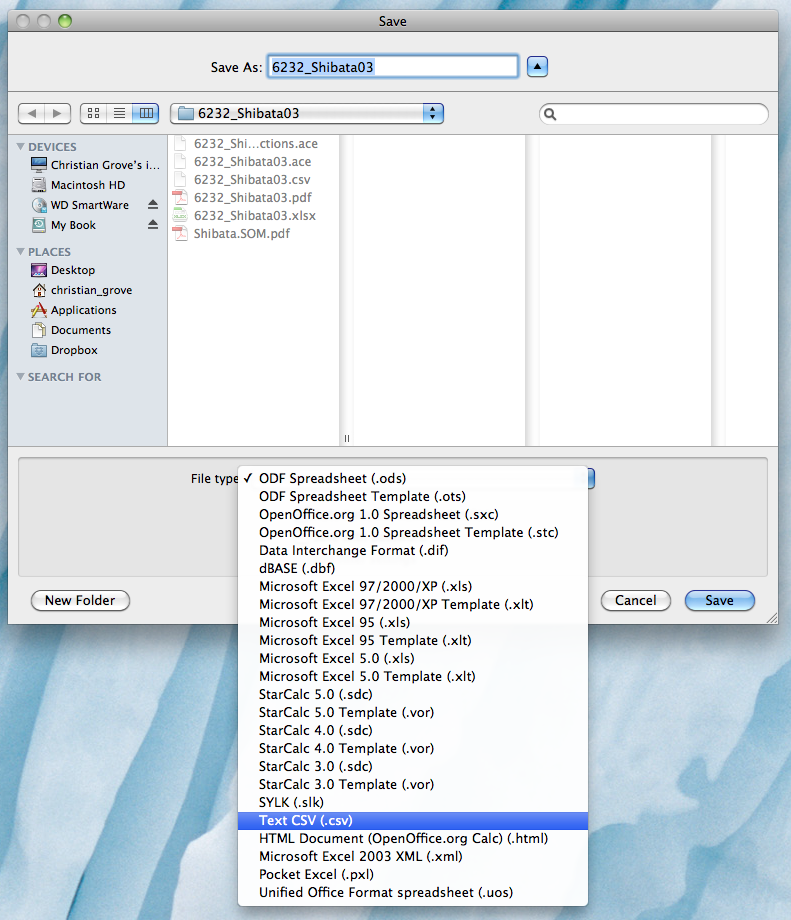
Once you have finished filling out the batch form and you are ready to submit it for processing into a *.ace file, you first need to convert it to a *.csv tab-delimited file. One option for doing this is to open the Excel document in Open Office, delete the yellow columns, and then "Save As..." and choose the "Text CSV (.csv)" option:
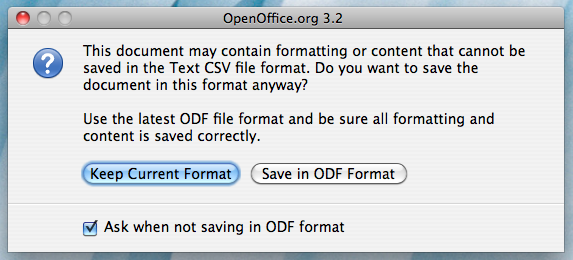
Once you click "Save", the following window may come up:
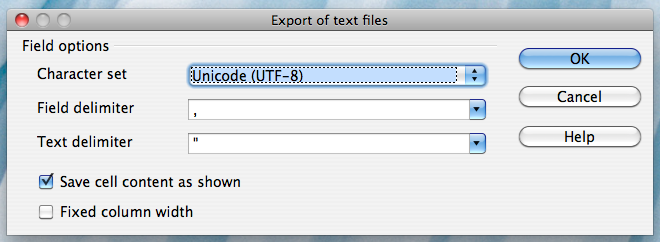
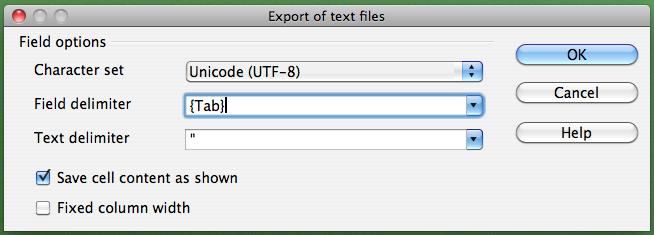
Click on "Keep Current Format". You will then be brought to the Open Office Export dialog box:
Non-Macintosh Export
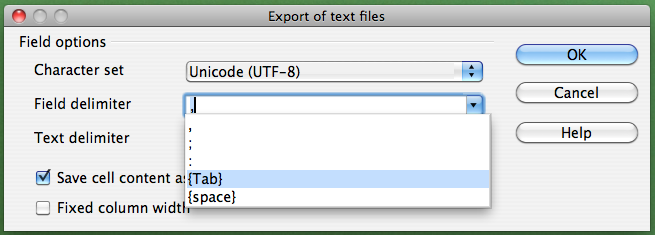
If you are not working on a Mac, click on the "Field delimiter" field, select "{Tab}", and then click "OK":
Macintosh Export
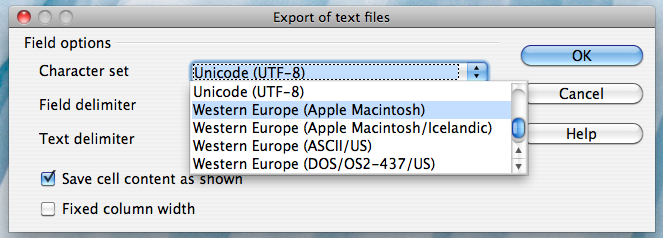
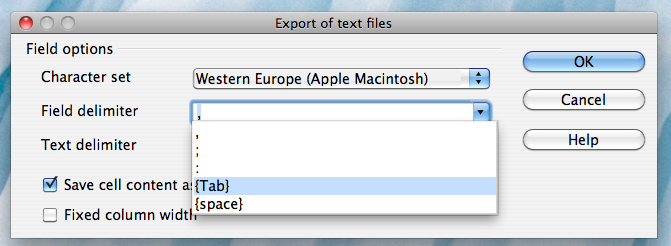
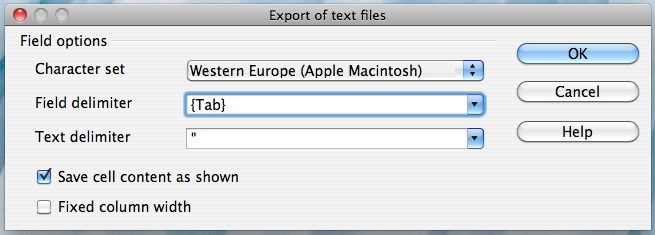
Alternatively, if you work on a Mac, select the "Character set" field in the dialog box and select "Western Europe (Apple Macintosh)". Next, select the "Field delimiter" field and select "{Tab}". Then click "OK"
This will save your *.csv file in the current directory (where the *.xls file is) or wherever you have specified.
Next, you will have to secure-copy your *.csv file to curation@elbrus.caltech.edu. Open a terminal, cd to your directory, and at the prompt type:
$ scp *.csv curation@elbrus.caltech.edu:~/RNAi/YourDirectory
(OR just "$ scp *.csv curation@elbrus:~/RNAi/YourDirectory" if you are on Caltech campus)
Enter the password.
Note: you will need to establish "YourDirectory" under the "RNAi" directory and place the batch form processing script into that directory.
Next, acquire secure-shell access to curation@elbrus:
$ ssh curation@elbrus
Enter the password.
Move to your directory:
$ cd RNAi/YourDirectory
Now run the Perl script with the following syntax:
$./rnai_batch_mapping.pl -i *.csv -t -o YourOutputName_Test.ace -e YourErrorFile_Test.txt
The "-i" option indicates your input *.csv file, the "-t" option indicates that you are running a test process of the *.csv file that will use temporary WB RNAi IDs (in case there are errors you don't want to assign new WBRNAi IDs), the "-o" option indicates the output *.ace file, and the "-e" indicates the error text file that will capture and report any errors that occur during processing.
The script will need to run for a few minutes (maybe several minutes depending on the size of you *.csv file). Once the script is done, check the error.txt file for any errors that may have occurred. If errors did occur, you will need to track down the source of the errors, fix them, and re-run the test processing.
The script generates two *.ace files: YourOutputName_Test.ace and RNAi_interaction.ace
Once your processing is free of errors, you may wish to test both *.ace files by reading them into ACEDB and ensuring that they read into the database with no errors.
Once your processing is free of errors, you will run the "real" processing as follows:
$./rnai_batch_mapping.pl -i *.csv -r -o YourOutputName.ace -e YourErrorFile.txt
Note that the only change is from using the "-t" option to the "-r" option. The "-r" option assigns real WB RNAi IDs to the RNAi experiments in the *.ace file.
Once the real processing runs smoothly and there are no errors, test the two *.ace files by reading them into ACEDB and making sure they read in without any errors. If both files read in without error, both files are ready to be submitted for upload to the database. Secure-copy the *.ace files to your local directory and archive them as needed.
Uploading the RNAi interactions .ACE file to the Interaction OA (Ontology Annotator)
Once you've generated you're RNAi interactions .ACE file and tested it by reading it into ACEDB (with no errors), you can upload the file to the Interactions OA. To do this, follow the instructions on the Gene Interaction Wiki Page
RNAi Paper Flagging and Processing of SVM "Low" Results and Tracking
We have been using SVM to flag papers that may contain RNAi data. These papers are given a probability score (High, Medium or Low) based on the SVM training set to reflect the possibility that they contain RNAi data to be curated. The High and Medium scoring papers are automatically added to the list of RNAi papers that are to be curated (on the Paper Editor) and the Lows are stored separately. The Low scoring papers must be manually checked by a curator and if the paper has "curatable" RNAi data, the curator must manually add that paper to the list. The Low scoring paper list is being checked approximately every 3 months (it is added to automatically every Monday at 2am based on SVM results). The following is the SOP for checking these papers and adding them to the list of RNAi papers to curate.
SVM Low's - SOP for checking papers
The file containing the papers with a low priority score (named "low") is stored on tazendra in the directory: /home/postgres/work/pgpopulation/svm/gary_rnai New papers are concatenated to this list. The curator checking the recently added papers must go back in the list to the most recent paper that has a "commented out" mark "//". This mark means that the list has been checked up to this point in the paper list....all papers after this mark need to be manually checked. When the curator is finished, they should "comment out" the last paper on the list so the next curator knows where to begin checking next time.
SVM Low's - SOP for adding papers to curation pipeline
After checking the "Low" papers, the ones that do conatin data must be added to the curation list. To do so go to the curator_first_pass.cgi on the web which is listed under "curation Forms" on the site map for tazendra. Add the appropriate number of the WBPaper id, for example just "00038308" without the WBPaper prefix, and click on the query button. Under "rnai" in the Gene Function section add the following comment in the curator box "SVM -LOW". When you hit "Flag" at the bottom of the page this paper will be added to the RNAi curation list and noted as an SVM with a low probability score.
SVM Low's - SOP for adding papers SVM Tracking
For tracking of SVM results (if False Positive, False Neg etc) a tool was created: http://XXXXXXXXX.caltech.edu/~postgres/cgi-bin/svm_results.cgi This web-based tool will automatically track papers that are positive based on our RNAi curation pipeline/checkout form and the OA; That is if a paper is on the checkout form and is curated, the data will be in the OA and marked as a True Positive. If the paper on the checkout list is a False Positive, the curator must note this on the paper editor (Flag False Positive) and the results will be forwarded to the SVM tracking form. http://tazendra.caltech.edu/~postgres/cgi-bin/paper_editor.cgi The SVM lows that do contain RNAi data are added to the checkout form by a curator (see above) and will be dealt with as any other paper on the list. The SVM lows that actually do not contain RNAi data must be entered manually onto the SVM tracking form as False Positives. To do this go to http://XXXXXXXXX.caltech.edu/~postgres/cgi-bin/svm_results.cgi Under "Enter Curator Results" Select data type/ select curator negative / add the list of negatives to / select comment "SVM positive - Curator negative"
RNAi OA
NOTE: Warning: RNAi objects may have multiple lines in the OA due to the nested phenotype data (Penetrance info, etc.)
TAB1
- pgid - the postgres ID - NOT DUMPED
- RNAi ID - rna_name - RNAi : - WBRNAiID #Note: This should be automatically assigned 8-digit padding. Assigned automatically based on highest value in rna_name table, if data is created without the OA, make sure to create a new OA line and change the name of the RNAi ID directly through postgres
- Paper - rna_paper - Reference - ?Paper (Ontology)
- Laboratory - rna_laboratory - Laboratory - ?Laboratory (Multi-ontology)
- Date - rna_date - Date - Text (YYYY-MM-DD)
- Curator - rna_curator - NOT DUMPED - Curator (Dropdown)
- PCR Product - rna_pcrproduct - PCR_product - ?PCR_product (Multi-ontology) #Note: EBI/Hinxton will map these
- DNA Text - rna_dnatext - DNA_text - Big text #Note: multiple Sequences should be separated by pipe "|" (caution: pressing <ENTER> may cause problems)
- Sequence - rna_sequence - Sequence - Text #Note: multiple DNA texts should be separated by pipe "|" (caution: pressing <ENTER> may cause problems). Not an ontology because there are over 2 million sequence objects in WS ; the gin_sequence only has those that have a WBGene associated.
- Strain - rna_strain - Strain - ?Strain (Ontology) - Already exists in Postgres; No files need to be made for this
- Genotype - rna_genotype - Genotype - Big text
- Treatment - rna_treatment - Treatment - Big text
- Life Stage - rna_lifestage - Life_stage - ?Life_stage (Ontology) - Already exists in Postgres; No files need to be made for this
- Temperature - rna_temperature - Temperature - Text (Integer)
- Delivery Method - rna_deliverymethod - Delivered_by - (Multi-dropdown) four choices: "Bacterial_feeding", "Injection", "Soaking", "Transgene_expression"
- Species - rna_species - Species - ?Species (Dropdown) #Note: Add Caenorhabditis_elegans to dropdown list added to species type to top of list
- Remark - rna_remark - Remark - Big text
TAB2
- From Genereg - rna_fromgenereg - NOT DUMPED - Toggle #Note: We would like a toggle here for Xiaodong to create new RNAi objects while performing Gene_regulation curation and flag them for an RNAi curator to complete (just like the "From RNAi" toggle in the Gene_regulation OA) Named field "From Gene Reg" to make it smaller, I'd prefer "From Genereg" or "From GeneReg", if you'd really rather have "From Gene Regulation" let me know and I'll change it. This is important before we start curation because the name of the field is the value that gets stored then the field is toggled on, so changing the field name later would mean changing the data in postgres as well -- J Sure, this sounds fine. We can keep it as small as you want; so, "From Genereg" is fine with me -- C
- NO DUMP - rna_nodump - NOT DUMPED - toggle - Would it be possible to add a 'NO DUMP' toggle here as well? I think we want to have Xiaodong turn that on when she creates a new object from Gene_regulation curation so that a partially curated object doesn't get dumped until after an RNAi curator gets to it -- C added, but you might want it next to the From Genereg field so that the curator doesn't need to switch to an extra tab just to click another toggle. I'm assuming that they won't be entering a lot of data, and that they'd do this fairly regularly. Yes, this toggle should be right next to (under) the "From Genereg" toggle -- C
- Phenotype Observed - rna_phenotype - Phenotype - ?Phenotype ID & name (Multi-ontology)
- Penetrance From - To - rna_penfromto - (in-line with Phenotype) Range - Text (Integer-space-Integer)
- Penetrance - rna_penetrance - (in-line with Phenotype) Penetrance - dropdown like in phenotype OA
- Heat Sensitive - rna_heatsens - (in-line with Phenotype) Temperature_sensitive - Toggle
- Cold Sensitive - rna_coldsens - (in-line with Phenotype) Temperature_sensitive - Toggle
- Quantity From - To - rna_quantfromto - (in-line with Phenotype) Quantity - Text (Integer-space-Integer)
- Quantity Description - rna_quantdesc - (in-line with Phenotype) Quantity_description - Big text
- Phenotype Remark - rna_phenremark - (in-line with Phenotype) Remark - Big Text -- Populate from Phenotype lines that have a Remark tag in #Phenotype_info following in .ace file.
- Molecule - rna_molecule - (in-line with Phenotype) Molecule - ?Molecule (Multi-ontology) This should be moved up inside the Phenotype subtags ? -- J Yes, that should do it -- C
- NOT - rna_phenotypenot - Phenotype_not_observed - NOT Toggle
TAB3
- Person Evidence - rna_person - Evidence Person_evidence - multiontology - for ?RNAi object's #Evidence This is the only dumped line that is giving an error. The dumper is making the line without the "Evidence" which is throwing errors. If the line reads "Evidence Person_evidence ..." then the *.ACE file will read in without any problems. -- C fixed, thanks -- J
- History Name - rna_historyname - History_name - Text #Note: this field is to accommodate older RNAi objects
- Movie - rna_movie - Movie - bigtext #Note: this field is to accommodate older RNAi objects You might want this to be a bigtext field, it stores URLs which are generally long, and some have multiples -- J OK, that sounds fine -- C
- Database - rna_database - Database - Text #Note: this field is to accommodate older RNAi objects I'm not sure how this will get displayed in this form. There are actually three inputs for this field in the RNAi model. There is 'Database', 'Database_field', and 'Accession_number'. It would be ideal to accommodate all three of these and display them in the OA. Does that make sense? -- C Enter data in this format, split multiple database lines with pipes if there are any :
- Phenobank2 Gene&RNAID GeneID=507328 | Phenobank3 Gene&RNAID GeneID=123456
- Expression Profile - rna_exprprofile - Expr_profile - Text #Note: this field is to accommodate older RNAi objects
NOT USING
- Evidence - postgres table and field not created, don't know what to parse into this field - Text #Note: this field is to accommodate older RNAi objects So there are only 8 RNAi objects that use the Evidence field, and they do so with 'Person_evidence'. In the OA, I think we should include: Person_evidence, Author_evidence, Curator_confirmed, Laboratory_evidence -- C We're only keeping Person_evidence for the #Evidence hash associated with the whole ?RNAi object -- C+J
- Penetrance Incomplete - rna_penincomplete - Toggle I think we talked about this, but in case we didn't phenotype OA uses a dropdown for the 4 penetrance types instead of separate toggle fields -- J OK, this sounds like a good idea. Let's just make it a dropdown list here as well -- C
- Penetrance Low - rna_penlow - Toggle
- Penetrance High - rna_penhigh - Toggle
- Penetrance Complete - rna_pencomplete - Toggle
- Method - rna_method - Method - Text
Notes
Fields for the tags "Predicted_gene", "Gene", "Transcript", "Pseudogene", "Homol_homol" and "Uniquely_mapped" will be omitted from the OA and populated in ACEDB after probe mapping to the genome at the EBI during the build process.
*.ACE Dumper Documentation
Each RNAi object will need to get dumped with "RNAi" in the "Method" tag
On tazendra, perl module is /home/postgres/work/citace_upload/rnai/get_rnai_ace.pm
Run with script /home/postgres/work/citace_upload/rnai/use_package.pl
Generates rnai.ace.<date> and err.out.<date> (to the day, so it will overwrite previous dumps from the same date) please check dumper and .ace file dumped -- J The dumper seems to be working OK since the dumped *.ACE file you generated is reading in OK, except for the "Evidence" line, as I comment about above in the "Tab3" section. The line currently dumps out as "Person_evidence ..." and it needs to dump out as "Evidence Person_evidence ...", otherwise the file throws errors when reading into ACEDB -- C Fixed, I thought only the innermost tag mattered, but I guess because there's no data in the RNAi part, just the tag, it needs to be there before the #Evidence part -- J I believe so, yes. -- C
Use_package.pl line comments:
'my $outfile = 'rnai.ace.' . $date;' 'my $errfile = 'err.out.' . $date;'
- These lines (above) specify the output; change as necessary.
'my ($all_entry, $long_text, $err_text) = &getRnai('all');'
'# my ($all_entry, $long_text, $err_text) = &getRnai('WBRNAi00008227');'
- The lines above specify what to dump out of the OA. 'all' dumps everything. If you want to dump specific objects, comment out the top line (add a '#') and uncomment the bottom line (remove '#') and specify the object name where 'WBRNAi00008227' is. Because of permissions, the script will need to be copied, modified, and then run from a permissible directory.
Molecule now converts the pgid stored in postgres with the molecule name in mop_name for that pgid -- J
<Chris needs to note the features of the script here>
Parsing WS RNAi into OA
There papers are not getting parsed into postgres tables for RNAi : WBPaper00004402, WBPaper00004403, WBPaper00004651, WBPaper00005654, WBPaper00006395, WBPaper00024497, WBPaper00025054, WBPaper00029258
/home/postgres/work/pgpopulation/rna_rnai/makeRnaiCuratorMappings.pl generates a mapping of RNAi objects to WBPerson curators based on WS231RNAiwithTimeStamp.ace
Parsing script is /home/postgres/work/pgpopulation/rna_rnai/populate_rnai.pl : populates postgres and generates /home/postgres/work/pgpopulation/rna_rnai/deletion.ace to remove all lines read into postgres + method lines which are not read into postgres but will get generated by dumper -- J OK, I just read Wen's RNAi objects from WS231 into an empty ACEDB (they read in fine), then I deleted their contents with the deletion.ace file (which read in fine), and then repopulated those RNAi objects with data dumped from the Sandbox RNAi OA (which read in fine). So, everything seems to be in working order. -- C 4/2/2012
Ignore all #Evidence in #Phenotype_info and in Remark tag of ?RNAi object.
To generate .ace deletion file always delete the Method line. Also delete anything getting parsed in.
These tags are not getting parsed into the OA : Author Gene Gene_regulation Homol_homol Interaction Predicted_gene Uniquely_mapped Correct -- C
These Phenotype subtags are not getting parsed into the OA : Curator_confirmed Molecule Paper_evidence . To see the Molecules not populated look on the sandbox at /home/postgres/work/pgpopulation/rna_rnai/phen_subtags_not_populated We do want these molecules in the OA. I guess I forgot/didn't realize that the molecules were coming from the "Affected_by" tag inside the #Phenotype_info hash. Will it be a problem to populate these molecules into the Molecule field in the OA? -- C It will probably be fine, since Molecule wasn't in the sub-section for Phenotype I thought it might have been a different type of molecule (like Laboratory is different from Laboratory_evidence, or something like that) You should move the field up into the Phenotype subtags in the wiki if I'm understanding that right. Just move it up to where you want it and I'll move it in the OA. I don't see the #Phenotype_info model in this wiki, and in the models.wrm I'm looking at there's no Affected_by tag, so I don't know how to parse it. The other #Phenotype_info was relatively straightforward because it was UNIQUE, but Molecule is multiontology so I'd need to make the parser more complicated to deal with that. Thing is that I don't see any data with multiple molecules. Do you know if there's any ? Does the field need to be a multiontology or is the data single value (so we can parse it that way) and we're making it multiontology in case it changes later ? -- J I've moved "Molecule" up into the Phenotype sub-tags and I've added the current (WS231) Phenotype_info model above, which includes the "Affected_by" and "Molecule" tags. I don't know if there are any annotations with multiple molecules, but even if there aren't any now, there's no reason there couldn't be one soon; therefore, I'd like to keep this multiontology, unless that is too much of a headache and we need to find another way around it. -- C There are many entries with multiple molecules, but they're parsed in. I missed this last night, but there's one molecule that isn't valid WBMol:00005097 /home/postgres/work/pgpopulation/rna_rnai/bad_molecule -- J OK, that bad molecule should be under the identifier D005467 instead of WBMol:00005097. Can we just replace each instance of WBMol:00005097 with D005467? -- C done -- Turns out this is the only object that has two pgids in the molecule OA right now, so I'm associating it to the lower of the two pgids : 3778 Molecule fields store in postgres the pgid of the corresponding molecule OA entry instead of the molecule name, because when the molecule is created sometimes it won't have a name. -- J
Is there only one Not phenotype in all the data ? (WBRNAi00085012) If so, it'd be a lot easier to ignore this, and after it's live add the phenotype manually through the OA. If not, let me know some example of other objects, because I can't find any to test for a parsing pattern. Is this the same as Phenotype_not_observed ? Hmmm, this is strange. We've tried to get rid of any "NOT" tags that remained in the RNAi objects. Wen had done this programmatically, but perhaps she missed this one. This is identical to Phenotype_not_observed -- C I'll wait until we resolve the NOT below, but probably just add it manually after everything else is transferred -- J resolved -- J
We're not keeping remark nor any other #Phenotype_info for the field Phenotype_not_observed right ? There's no place to put it Hmm, right; we would actually like to have remarks for Phenotype_not_observed. Can we add a "Phenotype_not_observed Remark" field? Given our earlier discussion, curators wouldn't put a "Phenotype" and a "Phenotype_not_observed" into the same row. -- C Ah, I didn't think it would work that way. I thought that positive phenotypes and negative phentoypes were completely different things so that you could add them on the same line the way you'd add a Curator and a Paper as unrelated data. If they're actually the same thing and should never be on the same line I'll parse it that way (it's easier), but then it would be easier to have a NOT toggle the way the phenotype OA works. If we do it this way the NOTs would be on separate lines depending on all the #Phenotype_info and we can keep the same Remark OA field for the dumper. It would just act like everything else but when the NOT toggle is on, it would use the Phenotype_not_observed in the .ace dump. We should talk before I parse this since it makes a difference -- J OK, that makes sense; let's handle Phenotype_not_observed like any other Phenotype and just add the NOT toggle. Then we can dump those out in the *.ACE file as "Phenotype_not_observed" plus any additional information captured in the #Phenotype_info hash. -- C done -- J
DNA_text is a double text field. I'm assuming that the first part is always going to be a string of atcg so will never have a space. In that case you could store the data in this field like qq(<atcg><space><next_thing>) but if you think you'd ever want spaces the better divider would be qq(<atcg>" "<next_thing") I didn't see anything else in the model that stores two values in a row besides the From-To fields, if there are any, let me know and we'll talk about them. I need to figure this out to know how to parse stuff. We talked about Database which I'm treating as a triple text field (instead of ontology values) and I think you don't have spaces in those values either, but if you do we should talk about it. I'm unclear about this; I thought that the "DNA_text" field would be a bigtext field that could accommodate multiple sequences <atcg> separated by pipes. The DNA_text field has two parts, the DNA and the probe. So you would separate different DNA_text lines with pipes, but need a different separator to separate DNA from probe -- J OK, I'm not sure what you mean by probe; unfortunately this term ("probe") gets thrown around a lot with different meanings. Can you give me an example? -- C Just got the word from the model above, but from what we talked about, let me know if you decide we don't what the probe part parsed in at all -- J As for the Database field, I thought we would enter it like this: Phenobank2 Gene&RNAID GeneID=507328, as discussed above. Is that not right? -- C Yes, the question is whether all those 5 things DNA, probe, database, database_field, and accession_number all have no spaces in their values. If they have spaces they need doublequotes around them, but if they'll never have spaces we can just have spaces in the OA when entering the text data -- J I see, I misunderstood. No, these values should never have spaces within themselves. -- C Cool, the current data doesn't have any spaces, so it's good. -- J
The OAs that have lifestage store the WBls:####### ID instead of the name, but the .ace file has names, so either there was some kind of translation, or more likely the data in there is wrong (someone should have switched it when the ?Life_stage class changed from text to IDs). I'm converting from names to IDs to store in postgres, but some names don't match the list of valid values in postgres in obo_name_lifestage. The list of RNAi objects and their Life_stage text is at /home/postgres/work/pgpopulation/rna_rnai/bad_lifestage OK, is it possible to correct these if I give you the proper names? "oocyte" should be "adult hermaphrodite" WBls:0000057, "Dauer" should be "dauer larva" WBls:0000032, "L3 larvae" should be "L3 larva" WBls:0000035, "young adult" should be "newly molted young adult hermaphrodite" WBls:0000063. I will have to look into "Mixed stages" and "L4-young adult". -- C I need the WBls IDs, I looked them up, added them earlier in the paragraph, and used them in the script. if you can find them for those last two that'd be good -- J OK for "Mixed stages" let's use "all stages" WBls:0000002 and for "L4-young adult" can we do two values? If we can I would use "L4 larva" WBls:0000038 and "newly molted young adult hermaphrodite" WBls:0000063. If only one value is allowed, just use "L4 larva" WBls:0000038. Thanks. -- C I think we could, but the Life_stage field is not multiontology do you want to switch it to multiontology or just go with WBls:0000038 (data currently parsed in with 38) OK, let's just leave it a single Ontology field (the model designates this as UNIQUE now anyway)-- C Also, why are we dumping Life_stage objects as names instead of WBls objects ? (probably talk to Wen about that) -- J Yeah, I have no idea about that. We'll have to ask Wen. -- C k, I'll leave as is for now -- J
Parsing the sequence data all 777 entries don't have a valid sequence in gin_sequence at /home/postgres/work/pgpopulation/rna_rnai/bad_sequence this might be because they all don't have WBGenes or something else. Is there a set that we should use like pcr_product ? Do you know why none of them match, is it the wrong tag or datatype in there ? The gin_sequence data is populated from genes that have a Corresponding_CDS or Corresponding_Transcript tag -- J Do the sequence objects need to be linked to a gene at the outset? That should be taken care of at the mapping stage. I've put the complete list (WS230) of sequence objects on tazendra at /home/acedb/chris/WS230_Sequence_objects.txt. -- C cool, I compared all the existing sequence entries, and they're all there, but that's not surprising since the RNAi sequences would create sequence objects in acedb by XREF, so it doesn't necessarily prove much. Because there's so many possible sequence objects (2 million) it'd probably be best to leave it as pipe-separated text the way that it's parsed in now, but we should talk about it -- J Yes, as we discussed, these will remain pipe-separated text objects -- C
There are a lot of pcr_products at /home/postgres/work/pgpopulation/rna_rnai/bad_pcrproduct but we might be able to get a lot more matches if we make it case insensitive. Does case matter ? No, case doesn't matter and I think you're right that that is probably the problem. -- C allowing upper and lower case in letters after the period, and they all match now -- J Great! -- C
There are a couple of bad papers at /home/postgres/work/pgpopulation/rna_rnai/bad_papers One paper is invalid linked to a lot of RNAi, the other has a typo lowercase p OK, it looks like WBPaper00013501 changed its name to WBPaper00024307, and, as you pointed out, the other is just a case typo. Can we just make the appropriate changes to these, or do we need to be more formal about it? -- C Hardcoded both exceptions, they're okay now -- J Excellent! Thanks! -- C
For all these problem objects if you want to fix them, we can edit the source .ace file, but if we ever switch the dataset to populate, we'd have to make all the changes in the new .ace file again. Probably easiest would be to tell me which objects to ignore and I can hardcode in not giving an error for those, not populating postgres with them, and keep track to populate through the OA when live. (if you want to keep track and change in the .ace file, that's easier for me, but in the past we've had to make dumps of .ace several times and used various sets so if it's the same with this datatype, it will be more of a pain for you I'd prefer, if possible, to make the changes directly to the .ace file(s) and thereby fix them permanently, unless there's a good reason not to do this. -- C You can sure. You shouldn't need to since I hardcoded those exceptions in the parser, so they'll go into postgres correctly -- J Great! Thanks! -- C
The .ace file has the species as "Caenorhabditis elegans", the value you wanted me to add to the OA (fits the other values's format) is "Caenorhabditis_elegans" so I'm converting from space in .ace to underscore for postgres. OK, I think leaving them both with a space is fine. I don't think there's any need for an underscore anywhere. -- C Well, all the other species have underscores, which is why I parsed it from space to underscore -- J OK, whatever is easiest. -- C
To sync to tazendra
- Create OA tables /home/postgres/work/pgpopulation/rna_rnai/create_rnai_tables.pl
- Create obo tables for pcr product data /home/postgres/work/pgpopulation/obo_oa_ontologies/20120322_pcrproduct/create_obo_tables.pl
- Populate obo tables for pcr product based on WS229_PCR_products.txt with /home/postgres/work/pgpopulation/obo_oa_ontologies/20120322_pcrproduct/populate_obo_pcrproduct.pl (takes 30 minutes, lots of lines)