Difference between revisions of "Ontology Annotator"
| Line 80: | Line 80: | ||
[[File:Google_Spreadsheet_Download_as_TSV_11-19-2013.png]] | [[File:Google_Spreadsheet_Download_as_TSV_11-19-2013.png]] | ||
| + | |||
| + | Even though the file menu suggests that the file will download as a "*.txt" file, it should download the spreadsheet in tab-delimited (TSV) file format which can be directly uploaded to the OA through the batch OA submission script. | ||
| + | |||
Back to [[Caltech documentation]] | Back to [[Caltech documentation]] | ||
[[Category:Curation]] | [[Category:Curation]] | ||
Revision as of 19:49, 19 November 2013
Contents
Description
The Ontology Annotator (OA) is a curation tool developed by WormBase for a variety of curation purposes including the curation of phenotypes, attaching GO terms to genes, genetic interactions, transgenes, free-text descriptions of genes, and several other data-types. The OA uses CGI, javascript and a postgreSQL database and is web-based, eliminating issues that may arise due to operating system differences and the need for an user to install other dependent software. The OA in many ways is similar to [Phenote] (phenote.org). The OA mainly consists of an Editor, where new data is entered or where pre-existing data can be queried and edited, a Data-Table for data review and a Term-Information panel where information about a term like IDs, synonyms etc., is displayed. The OA includes features like term autocomplete from pre-loaded ontologies, a fast AJAX loading of terms, ability to save/query to/from a postgreSQL database, duplicating data, editing several lines of data at once, and filtering the data. The display to some extent can be custom-organized as columns in the Data-Table can be sorted by dragging and the width of each data-column adjusted. For complex data-types with several fields, the OA allows a tabbed organization.
List of WormBase OA configurations:
- Antibody
- Concise description
- Disease
- Expression pattern
- Gene class
- Gene ontology
- Gene regulation
- Interaction
- Molecule
- Movie
- Phenotype
- Picture
- Process Term
- RNAi
- Topic
- Transgene
The OA uses :
- Perl CGI.
- Yahoo!'s YUI library and a local javascript file.
- PostgreSQL database backend (could probably be modified to other SQL databases).
- Apache webserver.
- Documentation for main CGI, javascript, and modules:OA docs
The above description can also be found here: Web-page for OA
Wish List
- Include dependencies wherever possible. For example, if making an IMP annotation for a given gene, have a gene-specific drop down menu of alleles or RNAi experiments for the WITH column. Or, if making an IGI annotation for the paper, have a drop down list of all genes mentioned in the paepr. Similarly, when entering a GO term, have the ontology (P, F, or C) get entered automatically. (From Curation Interface Meeting)
- Term information window - information should reflect where cursor is placed in the editor window, e.g. Paper should reflect paper info
Batch upload to OA from tab-delimited file
[New as of November 2013] A script has been written that will allow curators to upload data in bulk to the OA through the submission of a properly formatted tab-delimited (TSV) file. The script is located on Mangolassi/Tazendra at:
/home/postgres/public_html/cgi-bin/oa/scripts/populate_oa_tab_file/populate_oa_tab_file.pl
Usage
Enter (cd into) the directory with the script and run by entering:
./populate_oa_tab_file.pl mangolassi #### testfile
to enter into the Sandbox OA (on Mangolassi) where '####' is the curator's WBPerson ID, and 'testfile' is a test upload file to make sure everything is formatted properly
./populate_oa_tab_file.pl tazendra #### realfile
to enter into the Live OA (on Tazendra) where '####' is the curator's WBPerson ID, and 'realfile' is the real upload file that has been successfully tested on the sandbox (mangolassi)
Note: It is very important to test your batch upload file on the sandbox first, as there are many possible errors in formatting (mis-spelled names, IDs, etc.). Once uploaded to the OA on Tazendra, each entry will need to be edited manually, one-by-one if there are any mistakes.
OA's capable of accepting bulk upload
As of November 2013, the list of OA's that can accept bulk uploads via this method are as follows:
- Interaction
- Phenotype
- Process Term
- RNAi
- Topic
- Transgene
Tab-delimited (TSV) file format
It is important that the TSV file be formatted properly. Each column header must be a Postgres table name into which data will be uploaded. Each column header on a single form should be a Postgres table name for the same OA such that each row in the spreadsheet/TSV file will be a single entry (with a unique PGID) in the OA/Postgres.
Note: Every entry in a cell directly below a column header/Postgres table name MUST be entered EXACTLY as can be received by that table/field, i.e. if an 'ontology' or 'multiontology' field, a drop-down menu, or any other controlled vocabulary field, the entries must be formatted appropriately. If these data are not entered in the proper format, the data will enter the Postgres tables (incorrectly) but will not necessarily show up in the OA, making it difficult to track down erroneous data entries.
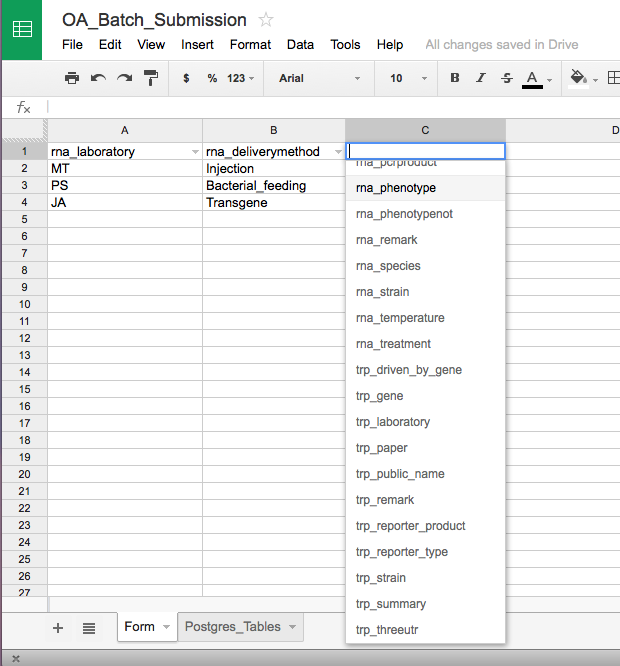
Any tab-delimited (TSV) file should work, but in order to standardize the submission process, a Google spreadsheet has been generated with drop down fields in each header row for selecting the correct Postgres table name:
https://docs.google.com/spreadsheet/ccc?key=0AgaLIpaBTJmSdEVoTXFLdHROOXQ1QzlkZ1VhYVpGMFE#gid=0
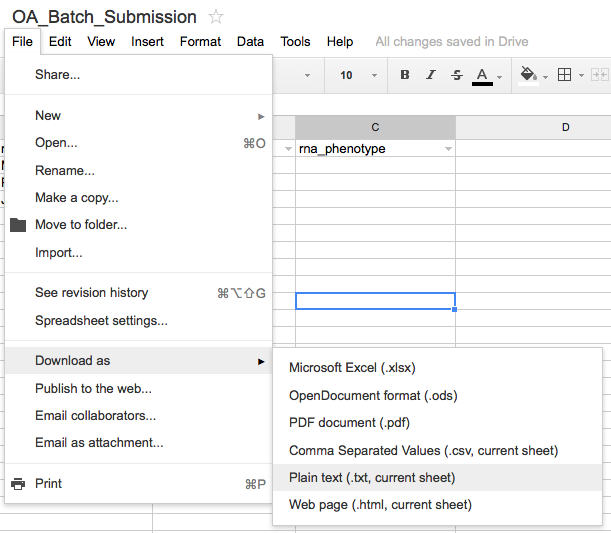
Above is a screenshot of the Google spreadsheet form. To download the form as a TSV file, select "Download As" from the "File" menu and select "Plain Text".
Even though the file menu suggests that the file will download as a "*.txt" file, it should download the spreadsheet in tab-delimited (TSV) file format which can be directly uploaded to the OA through the batch OA submission script.
Back to Caltech documentation