New 2012 Curation Status
Curation Status & Statistics Form (2012)
The sandbox/testing form can be found here
The CGI code is located on Tazendra/Mangolassi here:
/home/postgres/public_html/cgi-bin/curation_status.cgi
Contents
- 1 Pages of the Curation Status Form
- 2 Code Documentation
- 3 Abbreviations
- 4 Definitions
Pages of the Curation Status Form
Main Page
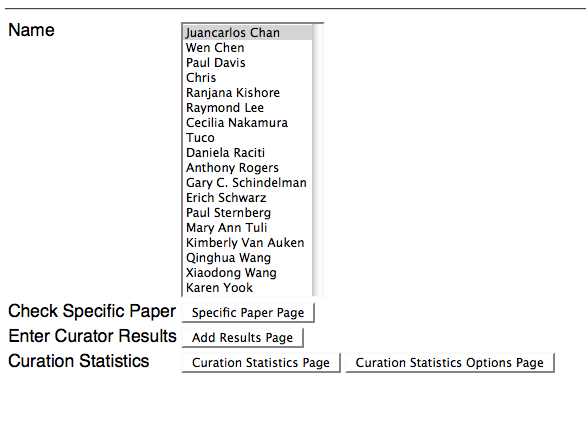
Above is a screenshot of the main page of the Curation Status Form. The user/curator is requested to identify who they wish to login as, and to select one of four options to continue:
1) Specific Paper Page - This is where the curator can specify one or more specific papers they wish to view curation status results for (see below).
2) Add Results Page - This is where the curator can add curation status results for one or more specific papers (see below).
3) Curation Statistics Page - This is where the curator can view all curation statistics for ALL datatypes and ALL flagging methods (see below).
4) Curation Statistics Options Page - As an alternative to viewing the curation statistics for ALL datatypes and ALL flagging methods (as with option #3 above), this is where the curator can specify which datatypes and flagging methods they would like to see curation statistics for (see below).
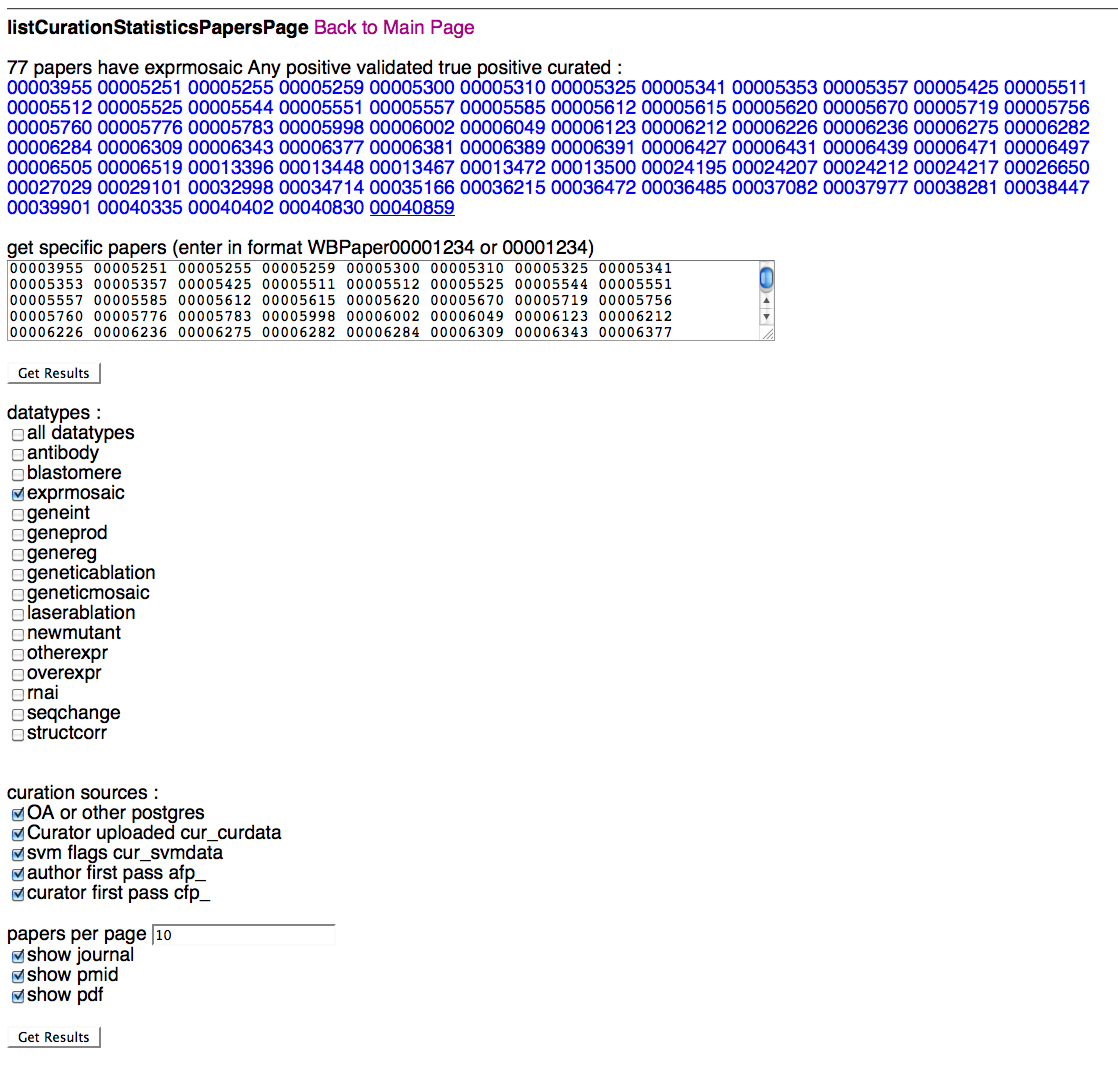
Specific Paper Page
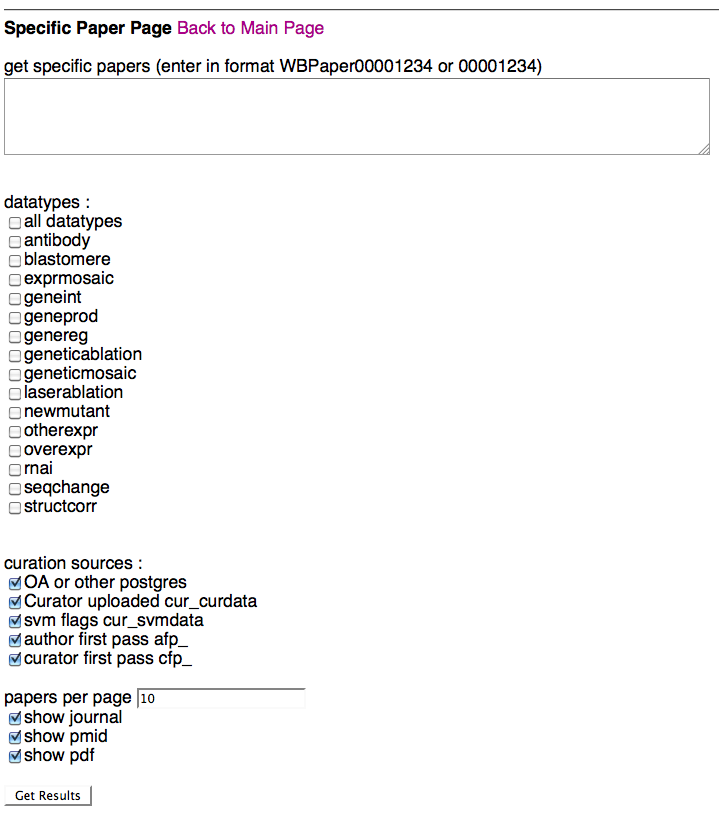
Above is a screenshot of the Specific Paper Page where a curator can specify which paper(s) they would like to view curation status results for. After typing/pasting in one or more WBPaper IDs in the paper entry field, the curator can specify which datatypes and flagging methods they would like to see results for. Note that selecting "all datatypes" will override any single datatype selections below. A curator can select what curation data sources they would like to see results for (i.e. Ontology Annotator and/or cur_curdata), flagging methods (SVM, AFP, CFP), the number of papers they would like to load at one time (default of 10), and whether they would like to see info (and links) for the PubMed ID (PMID), the PDF, and the paper's journal.
Once a curator clicks on "Get Results", they will be directed to the Detailed Results of Papers Page, where they can view the results of their query.
Papers can be listed as WBPaper### or simply as numbers, separated by spaces, commas, pipes, new lines, anything that is not a number. Any number that is entered will be considered a valid paper ID.
Add Results Page
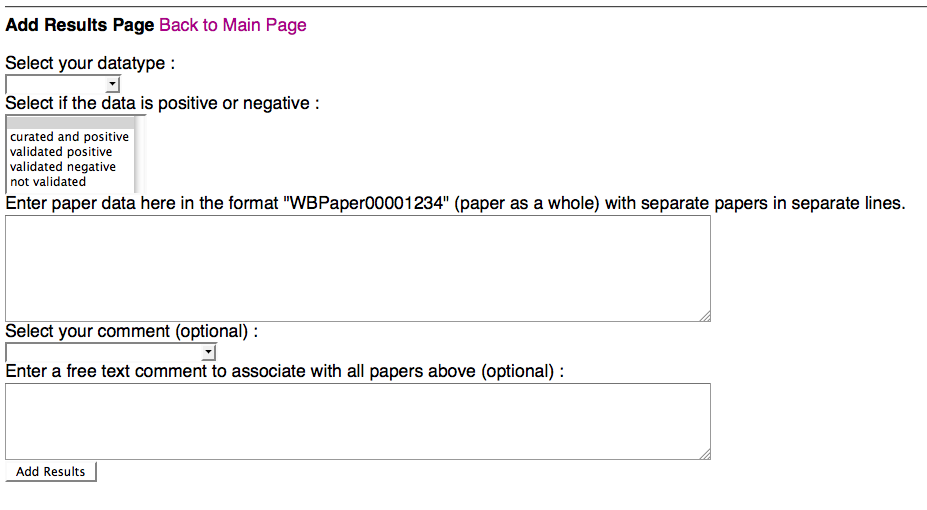
Above is a screenshot of the Add Results Page of the form, where a curator can add new curation status results for one or more papers that they specify. A curator must specify what datatype they wish to submit paper results for and must specify what the status is for the paper(s): curated and (hence) positive, validated postive (but not yet curated), validated negative, or (if they need to revert back to not validated, or blank, status) not validated. The curator must then also specify at least one paper for which to apply this curation status in the paper entry field. Multiple papers must be entered as WBPaper### format and each on a separate line.
Optionally, a curator can select a pre-made comment from a drop down menu and/or enter a free-text comment. Once the curator clicks "Add Results", they will be directed to a New Results Summary Page:
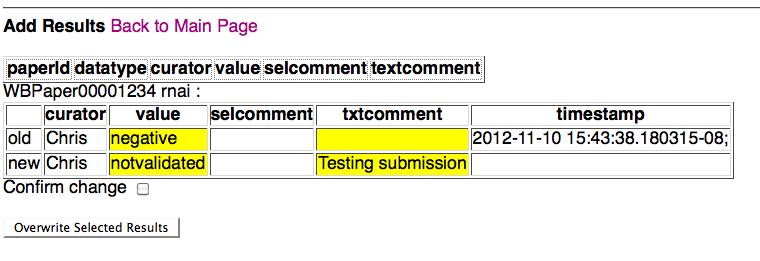
If the results are overwriting existing results, they will be directed to an Overwrite Confirmation Page:
at which point the curator can confirm the overwrite of the previous results for the indicated paper and datatype, or simply go the main page (or go back a page to make corrections/edits). Note that the fields for which data has changed are highlighted in yellow for easy viewing. If the curator confirms the overwrite by checking the confirmation check box and clicking on "Overwrite Selected Results", they will be directed to the Overwrite Confirmation Summary Page:
A link is provided to go back to the main page of the form.
Main Curation Statistics Page
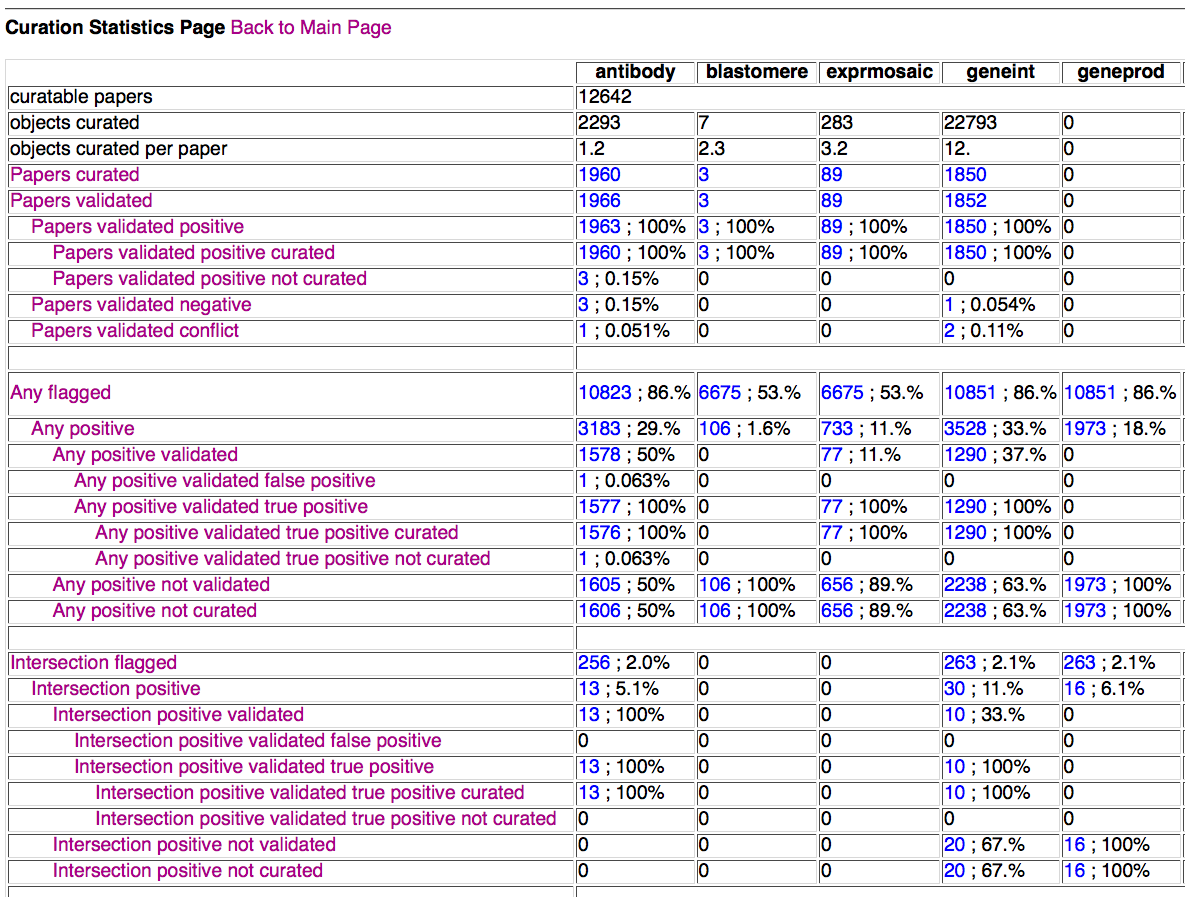
Above is a screenshot of a portion of the entire Curation Statistics table that a curator would be directed to from the main page of the form if they had clicked on the Curation Statistics Page button. Displayed at the top of the table are general paper statistics for a given datatype (datatypes indicated at the top of each column). Below that are statistics for papers that have been flagged (positive or negative) for the indicated datatype by ANY (at least one) flagging method. Below the "Any" statistics are the "Intersection" statistics, indicating papers flagged by ALL flagging methods for the indicated datatype. It should be emphasized here that "flagged" means processed by the flagging method, not necessarily flagged positive. Although not visible in the above screenshot, statistics for SVM results, AFP results, and CFP results are also included in this table.
The "Any", "Intersection", and individual flagging method sections of the table each follow a general template:
Flagged
Flagged Positive
Flagged Positive and Validated
Flagged Positive, Validated False Positive
Flagged Positive, Validated True Positive
Flagged Positive, Validated True Positive, Curated
Flagged Positive, Validated True Positive, Not Curated
Flagged Positive, Not Validated
Flagged Positive, Not Curated
and the individual flagging method sections additionally have a section for flagged negatives:
Flagged Negative
Flagged Negative and Validated
Flagged Negative, Validated True Negative
Flagged Negative, Validated False Negative
Flagged Negative, Validated False Negative, Curated
Flagged Negative, Validated False Negative, Not Curated
Flagged Negative, Not Validated
Flagged Negative, Not Curated
Each row title/header can be clicked on to bring up a small pop-up window with a brief description of what each title means. Each cell of the table has numbers indicating the number of papers that fit the criteria for that datatype and flag status, and the percentage (to two significant digits) that represents of a subset of some larger set. Each percentage is calculated, generally, as follows:
Flagged (% of curatable papers)
Flagged Positive (% flagged)
Flagged Positive and Validated (% flagged positive)
Flagged Positive, Validated False Positive (% flagged positive and validated)
Flagged Positive, Validated True Positive (% flagged positive and validated)
Flagged Positive, Validated True Positive, Curated (% flagged positive and validated true positive)
Flagged Positive, Validated True Positive, Not Curated (% flagged positive and validated true positive)
Flagged Positive, Not Validated (% flagged positive)
Flagged Positive, Not Curated (% flagged positive)
Each cell number (aside from the top three rows) is also a hyperlink to the Prepopulated Specific Papers Page, listing the paper IDs for each paper in the list, as well as providing options for the view of each of those papers in the Detailed Results of Papers Page.
Curation Statistics Page Display Info
The title/headers for each row are displayed at the left AND right sides of the table, to enable easier viewing when there are several datatypes being viewed at once. If the number of datatypes is restricted via the Curation Statistics Options Page, the titles/headers for each row will only display on both left and right sides of the table if more than six datatypes are selected for viewing (this was done to avoid overcrowding of the page when six or fewer datatypes/columns were visible).
The row-title column (leftmost column and, when more than six datatypes are visible, the rightmost column) are set to display at a fixed width of 600 pixels to allow all titles to fit on a single line. All other columns (datatype columns) are set to display at a fixed width of 120 pixels.
Curation Statistics Options Page
Above is a screenshot of the Curation Statistics Options Page, where a curator can specify what flagging methods and datatypes they would like to see curation statistics for. (Note 1) Loading a table with fewer datatypes or flagging methods is often much faster than loading the entire table. Whereas loading the entire table (with ALL datatypes and ALL flagging methods) takes roughly 23 seconds to load, loading a table with ALL flagging methods but ONE datatype will usually only take 1-3 seconds to load. (Note 2) If 6 or fewer datatypes are requested, the row titles/headers will only appear on the left side of the table; if more than 6 datatypes are viewed at once, the row titles/headers will appear on both the left AND the right sides of the table. (Note 3) The "Any" and "Intersection" rows of the resulting table will only show results for the flagging methods that you have selected to view. If only one flagging method is selected to view, the "Any" and "Intersection" rows will be identical to each other and to the "flagged positive" rows for the single flagging method.
Prepopulated Specific Papers Page
Above is a screenshot of the Prepopulated Specific Papers Page. Curators are directed here from any Curation Statistics table via the hyperlinked numbers in the statistics table. The entire list of paper IDs (that fit the criteria indicated in the table row/column of the statistics table) is listed as hyperlinks to the individual paper results (on the Detailed Results of Papers Page), as well as in the search box. Note that this page is identical to the Specific Paper Page, except that paper IDs are already pre-populated into the form from the statistics table.
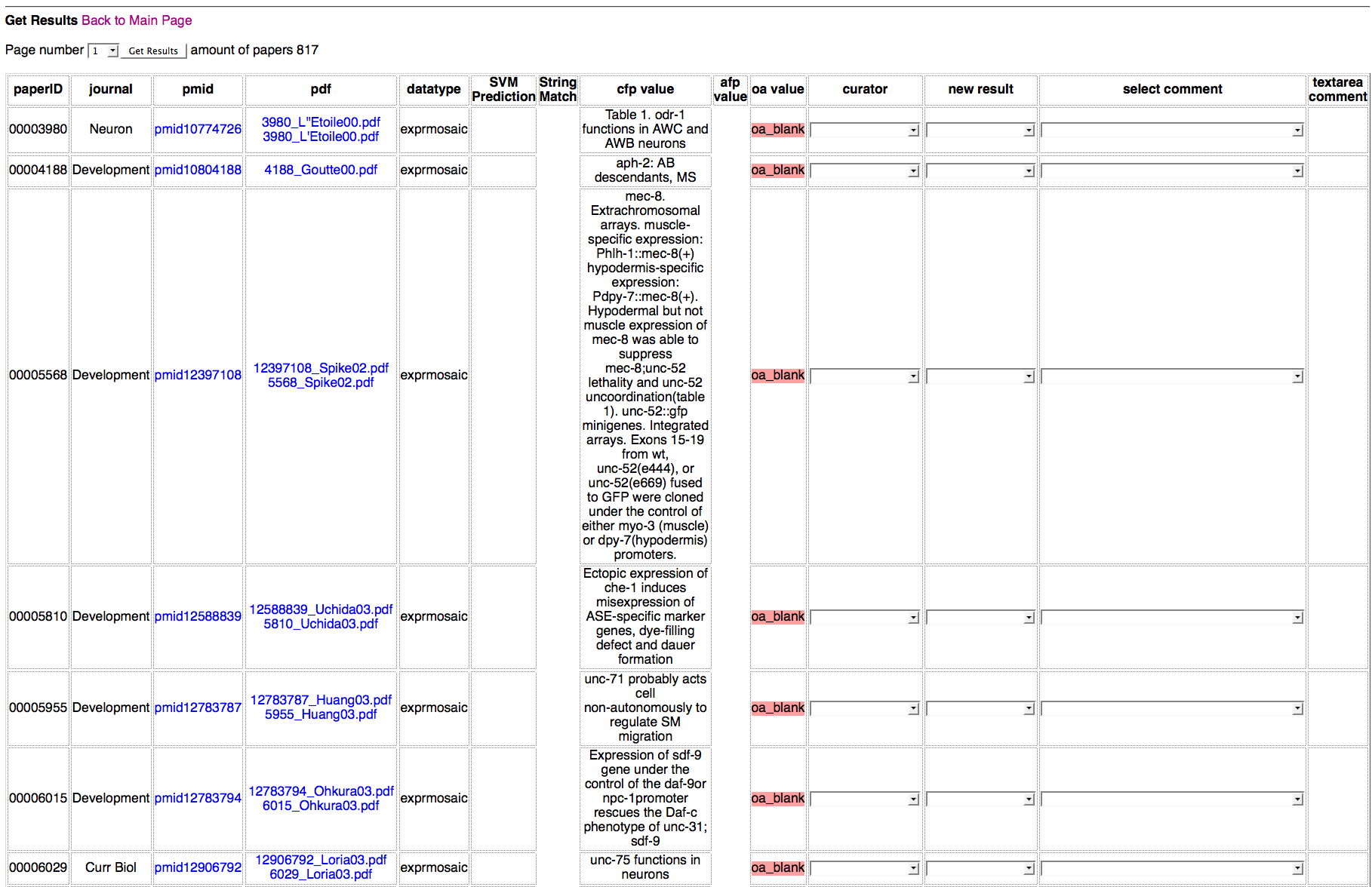
Detailed Results of Papers Page
Above is a screenshot of the Detailed Results of Papers Page, where curators can view and edit the curation status of individual papers as well as view the the flagging results of each flagging method for individual papers.
When ALL columns are visible:
the first column displays the WormBase paper ID (WBPaperID#);
the second column displays the name of the journal for the publication;
the third column lists the PubMed ID (PMID) and links out to the PubMed webpage for this article;
the fourth column displays the name of the PDF file stored locally on Tazendra with a hyperlink to the PDF file of the article in our local PDF archives;
the fifth column displays the datatype for that row (note that multiple datatypes may be displayed per paper, on separate rows);
the sixth, seventh, and eighth columns display the results of the flagging methods SVM, CFP, and AFP, respectively;
the ninth column indicates the status of the paper in the Ontology Annotator (OA) for that datatype indicating "oa_blank" if the paper does not exist in the respective OA, or "curated" if the paper does exist in the respective OA, indicating that it has been curated (or at least partially curated);
the tenth column provides a drop-down menu to select a curator (selecting a curator is only necessary if overwriting/changing the existing curator; the form recognizes what curator is logged in and automatically populates this field with the correct (logged in) curator if this field is blank);
the eleventh column provides a drop-down menu to select the "new result" for the paper, indicating whether it is "curated and positive", "validated positive", "validated negative", or "not validated" ("not validated" only needs to be selected when reverting back from "curated and positive", "validated positive", or "validated negative" entries that may have been entered accidentally)
the twelfth column provides the drop-down menu of standard, premade comments
the thirteenth (and final) column is a free-text area where a curator can write in any pertinent notes about the curation status of this paper-datatype pair
If any new results for a paper are entered (in columns 10-13), the curator must click on the "Submit New Results" button at the bottom of the screen, at which point they will either be directed to the New Results Summary page or to the Overwrite Confirmation Page, as shown above in the Wiki section describing the Add Results Page. Note that in order for new paper result submissions to take effect, there must be a value in the "new results" column. Otherwise any comments (premade or free-text) will not be registered with the paper.
Code Documentation
Below is the documentation for the form's code, located on Tazendra (when live) or Mangolassi (sandbox):
/home/postgres/public_html/cgi-bin/curation_status.cgi
Add Results Page: Loading Page and Processing Input
The following code is responsible for printing the Add Results Page:
sub printAddResultsPage {
&printAddSection('', '', '', '', '', '', '');
} # sub printAddResultsPage
This code defines the printAddResultsPage subroutine which in turn calls upon the printAddSection subroutine (see below), passing it empty strings, declaring empty/initialized values for the curator ($twonumForm), datatype ($datatypeForm), validation result ($donposnegForm), paper list ($paperResultsForm), premade comment ($selcommentForm), and free-text comment ($txtcommentForm). Once data is submitted, these variables acquire values and are reported in the event of an error.
The following code defines the printAddSection subroutine mentioned above, which adds the form components for entering datatype, validation status, paper IDs, premade comments, and free-text comments:
sub printAddSection {
my ($twonumForm, $datatypeForm, $donposnegForm, $paperResultsForm, $selcommentForm, $txtcommentForm) = @_;
my $selected = '';
&printFormOpen();
&printHiddenCurator();
print qq(Select your datatype :<br/>);
print qq(<select name="select_datatype">);
print qq(<option value="" ></option>\n);
foreach my $datatype (keys %datatypes) {
if ($datatype eq $datatypeForm) { $selected = qq(selected="selected"); } else { $selected = ''; }
print qq(<option value="$datatype" $selected>$datatype</option>\n); }
print qq(</select><br/>);
print qq(Select if the data is positive or negative :<br/>);
my $select_size = scalar keys %donPosNegOptions;
print qq(<select name="select_donposneg" size="$select_size">);
foreach my $donposnegValue (keys %donPosNegOptions) {
if ($donposnegForm eq $donposnegValue) { $selected = qq(selected="selected"); } else { $selected = ''; }
print qq(<option value="$donposnegValue" $selected>$donPosNegOptions{$donposnegValue}</option>\n); }
print qq(</select><br/>);
print qq(Enter paper data here in the format "WBPaper00001234" (paper as a whole) with separate papers in separate lines.<br/>);
print qq(<textarea name="textarea_paper_results" rows="6" cols="80">$paperResultsForm</textarea><br/>\n);
print qq(Select your comment (optional) :<br/>);
print qq(<select name="select_comment">);
print qq(<option value="" ></option>\n);
foreach my $comment (keys %premadeComments) {
if ($comment eq $selcommentForm) { $selected = qq(selected="selected"); } else { $selected = ''; }
print qq(<option value="$comment" $selected>$premadeComments{$comment}</option>\n); }
print qq(</select><br/>);
print qq(Enter a free text comment to associate with all papers above (optional) :<br/>);
print qq(<textarea rows="4" cols="80" name="textarea_comment">$txtcommentForm</textarea><br/>);
print qq(<input type="submit" name="action" value="Add Results"><br/>\n);
&printFormClose();
} # sub printAddSection
When a curator clicks on "Add Results" on the Add Results Page, the following code will process the curator's input, catching errors when they arise:
sub addResults {
&printFormOpen();
&printHiddenCurator();
my $errorData = '';
my %papersToAdd;
my $twonum = $curator;
($oop, my $datatype) = &getHtmlVar($query, "select_datatype");
unless ($datatype) { $errorData .= "Error : Need to select a datatype.<br/>\n"; }
($oop, my $donposneg) = &getHtmlVar($query, "select_donposneg");
unless ($donposneg) { $errorData .= "Error : Need to select whether result is curated, validated positive, or validated negative.<br/>\n"; }
($oop, my $paperResults) = &getHtmlVar($query, "textarea_paper_results");
if ($paperResults) {
my @lines = split/\r\n/, $paperResults;
foreach my $line (@lines) {
if ($line =~ m/^WBPaper(\S+)$/) { $papersToAdd{$1}++; }
else { $errorData .= qq(Error bad line : ${line}<br/>\n); }
} } # foreach my $line (@lines)
else { $errorData .= "Error : Need to enter at least one paper.<br/>\n"; }
($oop, my $selcomment) = &getHtmlVar($query, "select_comment");
($oop, my $txtcomment) = &getHtmlVar($query, "textarea_comment");
if ($errorData) { # problem with data, do not allow creation of any data, show form again
print "$errorData<br />\n";
printAddSection($twonum, $datatype, $donposneg, $paperResults, $selcomment, $txtcomment); }
else { # all data is okay, enter data.
my $joinkeys = join"','", sort keys %papersToAdd;
my ($pgDataRef) = &getPgDataForJoinkeys($joinkeys, $datatype);
my %pgData = %$pgDataRef;
my @data; my @duplicateData;
foreach my $joinkey (sort keys %papersToAdd) {
my @line;
push @line, $joinkey;
push @line, $datatype;
push @line, $twonum;
push @line, $donposneg;
push @line, $selcomment;
push @line, $txtcomment;
if ($pgData{$joinkey}{$datatype}) { push @duplicateData, \@line; }
else { push @data, \@line; }
} # foreach my $joinkey (sort keys %papersToAdd)
&processResultDataDuplicateData(\@data, \@duplicateData, \%pgData);
} # else # if ($errorData)
&printFormClose();
} # sub addResults
This code (above) will check to ensure the following:
1) A datatype has been selected
2) A validation status has been entered
3) At least one paper ID has been submitted
4) There is only one paper ID per line
5) The paper IDs entered are in the format 'WBPaper########'
Any exceptions to these will result in an error message printed to the screen, in addition to reprinting the screen with the submitted values in there respective fields.
If no errors are found, the script will continue by calling on the getPgDataForJoinkeys subroutine (to query Postgres for the cur_curdata associated with each paper-datatype pair; see below) and writing the new input values to the appropriate paper-datatype pairs.
sub getPgDataForJoinkeys {
my ($joinkeys, $datatype) = @_;
my %pgData;
$result = $dbh->prepare( "SELECT * FROM cur_curdata WHERE cur_datatype = '$datatype' AND cur_paper IN ('$joinkeys')" );
$result->execute() or die "Cannot prepare statement: $DBI::errstr\n";
while (my @row = $result->fetchrow) {
$pgData{$row[0]}{$row[1]}{curator} = $row[2];
$pgData{$row[0]}{$row[1]}{donposneg} = $row[3];
$pgData{$row[0]}{$row[1]}{selcomment} = $row[4];
$pgData{$row[0]}{$row[1]}{txtcomment} = $row[5];
$pgData{$row[0]}{$row[1]}{timestamp} = $row[6]; }
return \%pgData;
} # sub getPgDataForJoinkeys
The code will then run the processResultDataDuplicateData subroutine (see below) to print the results to the screen as the New Results Summary Page (for new data) and/or handle the overwrite confirmation when data needs to be overwritten, generating the Overwrite Confirmation Page (see Add Results Page section above).
sub processResultDataDuplicateData {
my ($dataRef, $duplicateDataRef, $pgDataRef) = @_;
my @data = @$dataRef;
my @duplicateData = @$duplicateDataRef;
my %pgData = %$pgDataRef;
print qq(<table border="1">\n);
print qq(<tr>${thDot}paperId</td>${thDot}datatype</td>${thDot}curator</td>${thDot}value</td>${thDot}selcomment</td>${thDot}textcomment</td></tr>\n);
foreach my $lineRef (@data) {
my @line = @$lineRef;
foreach (@line) { unless ($_) { $_ = ''; } } # initialize values if none are there
my $pgvalues = join"','", @line;
my @pgcommands = ();
my $pgcommand = "INSERT INTO cur_curdata VALUES ('$pgvalues');";
push @pgcommands, $pgcommand;
$pgcommand = "INSERT INTO cur_curdata_hst VALUES ('$pgvalues');";
push @pgcommands, $pgcommand;
foreach my $pgcommand (@pgcommands) {
print qq($pgcommand<br/>\n);
# UNCOMMENT TO POPULATE
$dbh->do( $pgcommand );
}
my $trData = join"</td>$tdDot", @line;
print qq(<tr>${tdDot}$trData</td></tr>\n);
} # foreach my $lineRef (@data)
print qq(</table>\n);
if (scalar @data > 0) { print "results added<br />\n"; }
my $overwriteCount = 0;
foreach my $lineRef (@duplicateData) { # for data already in postgres, add option to overwrite
my @line = @$lineRef;
foreach (@line) { unless ($_) { $_ = ''; } } # initialize values if none are there
my ( $joinkey, $datatype, $twonum, $donposneg, $selcomment, $txtcomment ) = @line;
my ( $curatorPg, $curatorPgName, $donposnegPg, $selcommentPg, $selcommentPgText, $txtcommentPg, $timestampPg ) = ( ' ', ' ', ' ', ' ', ' ', ' ', ' ' );
my ( $curatorFm, $curatorFmName, $donposnegFm, $selcommentFm, $selcommentFmText, $txtcommentFm, $timestampFm ) = ( ' ', ' ', ' ', ' ', ' ', ' ', '<td> </td>' );
if ( $pgData{$joinkey}{$datatype}{curator} ) { $curatorPg = $pgData{$joinkey}{$datatype}{curator}; $curatorPgName = $curators{$curatorPg}; }
if ( $pgData{$joinkey}{$datatype}{donposneg} ) { $donposnegPg = $pgData{$joinkey}{$datatype}{donposneg}; }
if ( $pgData{$joinkey}{$datatype}{selcomment} ) { $selcommentPg = $pgData{$joinkey}{$datatype}{selcomment}; $selcommentPgText = $premadeComments{$selcommentPg}; }
if ( $pgData{$joinkey}{$datatype}{txtcomment} ) { $txtcommentPg = $pgData{$joinkey}{$datatype}{txtcomment}; }
if ( $pgData{$joinkey}{$datatype}{timestamp} ) { $timestampPg = "<td>$pgData{$joinkey}{$datatype}{timestamp};</td>" }
if ( $twonum ) { $curatorFm = $twonum;
if ( $curators{$curatorFm} ) { $curatorFmName = $curators{$curatorFm}; } }
if ( $donposneg ) { $donposnegFm = $donposneg; }
if ( $selcomment ) { $selcommentFm = $selcomment;
if ( $premadeComments{$selcommentFm} ) { $selcommentFmText = $premadeComments{$selcommentFm}; } }
if ( $txtcomment ) { $txtcommentFm = $txtcomment; }
my $isDifferent = 0; # if any of the non-key values has changed, show option to overwrite
if ($curatorFmName ne $curatorPgName) {
$isDifferent++;
$curatorFmName = '<td style="background-color:yellow">' . $curatorFmName . '</td>';
$curatorPgName = '<td style="background-color:yellow">' . $curatorPgName . '</td>'; }
else {
$curatorFmName = '<td>' . $curatorFmName . '</td>';
$curatorPgName = '<td>' . $curatorPgName . '</td>'; }
if ($donposnegFm ne $donposnegPg) {
$isDifferent++;
$donposnegFm = '<td style="background-color:yellow">' . $donposnegFm . '</td>';
$donposnegPg = '<td style="background-color:yellow">' . $donposnegPg . '</td>'; }
else {
$donposnegFm = '<td>' . $donposnegFm . '</td>';
$donposnegPg = '<td>' . $donposnegPg . '</td>'; }
if ($selcommentFmText ne $selcommentPgText) {
$isDifferent++;
$selcommentFmText = '<td style="background-color:yellow">' . $selcommentFmText . '</td>';
$selcommentPgText = '<td style="background-color:yellow">' . $selcommentPgText . '</td>'; }
else {
$selcommentFmText = '<td>' . $selcommentFmText . '</td>';
$selcommentPgText = '<td>' . $selcommentPgText . '</td>'; }
if ($txtcommentFm ne $txtcommentPg) {
$isDifferent++;
$txtcommentFm = '<td style="background-color:yellow">' . $txtcommentFm . '</td>';
$txtcommentPg = '<td style="background-color:yellow">' . $txtcommentPg . '</td>'; }
else {
$txtcommentFm = '<td>' . $txtcommentFm . '</td>';
$txtcommentPg = '<td>' . $txtcommentPg . '</td>'; }
next unless ($isDifferent > 0);
$overwriteCount++;
print qq(<input type="hidden" name="joinkey_$overwriteCount" value="$joinkey" >);
print qq(<input type="hidden" name="datatype_$overwriteCount" value="$datatype" >);
print qq(<input type="hidden" name="twonum_$overwriteCount" value="$twonum" >);
print qq(<input type="hidden" name="donposneg_$overwriteCount" value="$donposneg" >);
print qq(<input type="hidden" name="selcomment_$overwriteCount" value="$selcomment" >);
print qq(<input type="hidden" name="txtcomment_$overwriteCount" value="$txtcomment" >);
print qq(WBPaper$joinkey $datatype : <br/>\n);
print qq(<table border="1">\n);
print qq(<tr><th> </th><th>curator</th><th>value</th><th>selcomment</th><th>txtcomment</th><th>timestamp</th></tr>);
print qq(<tr><td>old</td>${curatorPgName}${donposnegPg}${selcommentPgText}${txtcommentPg}${timestampPg}</tr>\n);
print qq(<tr><td>new</td>${curatorFmName}${donposnegFm}${selcommentFmText}${txtcommentFm}${timestampFm}</tr>\n);
print qq(</table>\n);
print qq(Confirm change <input type="checkbox" name="checkbox_$overwriteCount" value="overwrite"><br/><br/>\n);
} # foreach my $lineRef (@data)
if ($overwriteCount > 0) {
print qq(<input type="hidden" name="overwrite_count" value="$overwriteCount">);
print qq(<input type="submit" name="action" value="Overwrite Selected Results"><br/>\n); }
} # sub processResultDataDuplicateData
Once the curator has confirmed the overwrite of the relevant results, the overwriteSelectedResults subroutine (below) will run to officially overwrite the data in the Postgres cur_curdata table.
sub overwriteSelectedResults {
($oop, my $overwriteCount) = &getHtmlVar($query, "overwrite_count");
my @pgcommands;
for my $i (1 .. $overwriteCount) {
($oop, my $overwrite) = &getHtmlVar($query, "checkbox_$i");
next unless ($overwrite eq 'overwrite');
($oop, my $joinkey ) = &getHtmlVar($query, "joinkey_$i" );
($oop, my $datatype ) = &getHtmlVar($query, "datatype_$i" );
($oop, my $twonum ) = &getHtmlVar($query, "twonum_$i" );
($oop, my $donposneg ) = &getHtmlVar($query, "donposneg_$i" );
($oop, my $selcomment ) = &getHtmlVar($query, "selcomment_$i" );
($oop, my $txtcomment ) = &getHtmlVar($query, "txtcomment_$i" );
unless ($donposneg) { $donposneg = ''; } unless ($selcomment) { $selcomment = ''; } unless ($txtcomment) { $txtcomment = ''; }
push @pgcommands, qq(DELETE FROM cur_curdata WHERE cur_paper = '$joinkey' AND cur_datatype = '$datatype' AND cur_curator = '$twonum');
push @pgcommands, qq(INSERT INTO cur_curdata VALUES ('$joinkey', '$datatype', '$twonum', '$donposneg', '$selcomment', '$txtcomment'));
push @pgcommands, qq(INSERT INTO cur_curdata_hst VALUES ('$joinkey', '$datatype', '$twonum', '$donposneg', '$selcomment', '$txtcomment'));
} # for my $i (1 .. $overwriteCount)
foreach my $pgcommand (@pgcommands) {
print "$pgcommand<br />\n";
# UNCOMMENT TO POPULATE
$dbh->do( $pgcommand );
} # foreach my $pgcommand (@pgcommands)
} # sub overwriteSelectedResults
Get Results (Detailed Results of Papers)
The following code is for the getResults subroutine which displays the Detailed Results of Papers Page after receiving input from the curator about what to display (from a Specific Paper Page or Prepopulated Specific Paper Page submission).
The first section of code collects all curatable papers, processes the checkbox input for the various datatypes, and processes the paper IDs that were submitted in the paper field.
sub getResults {
&printFormOpen();
&printHiddenCurator();
&populateCuratablePapers(); # assume for now that we only care about curatable papers
($oop, my $all_datatypes_checkbox) = &getHtmlVar($query, "checkbox_all_datatypes");
unless ($all_datatypes_checkbox) { $all_datatypes_checkbox = ''; }
foreach my $datatype (keys %datatypes) {
($oop, my $chosen) = &getHtmlVar($query, "checkbox_$datatype");
unless ($chosen) { $chosen = ''; }
if ($all_datatypes_checkbox eq 'all') { $chosen = $datatype; } # if all datatypes checkbox was selected, set that datatype's chosen to that datatype
print qq(<input type="hidden" name="checkbox_$datatype" value="$chosen">\n);
if ($chosen) { $chosenDatatypes{$chosen}++; }
} # foreach my $datatype (keys %datatypes)
($oop, my $specificPapers) = &getHtmlVar($query, "specific_papers");
if ($specificPapers) { my (@joinkeys) = $specificPapers =~ m/(\d+)/g; foreach (@joinkeys) { $chosenPapers{$_}++; } }
else { $chosenPapers{'all'}++; }
print qq(<input type="hidden" name="specific_papers" value="$specificPapers">\n);
The next section of code populates cur_curdata for the respective paper-datatype pairs given the input from above. The code takes into account whether to show OA data and what flagging methods to display for each paper-datatype pair, to prepare them for display. Additionally, the code is now processing how many papers to display per page, setting the default page (if multiple pages of results) to see page "0", and determining whether or not the curator wishes to see the journal, PMID, and/or PDF links for each paper.
&populateCurCurData(); # always show curator values since they have to be editable
($oop, my $displayOa) = &getHtmlVar($query, "checkbox_oa"); unless ($displayOa) { $displayOa = ''; }
($oop, my $displayCfp) = &getHtmlVar($query, "checkbox_cfp"); unless ($displayCfp) { $displayCfp = ''; }
($oop, my $displayAfp) = &getHtmlVar($query, "checkbox_afp"); unless ($displayAfp) { $displayAfp = ''; }
($oop, my $displaySvm) = &getHtmlVar($query, "checkbox_svm"); unless ($displaySvm) { $displaySvm = ''; }
print qq(<input type="hidden" name="checkbox_oa" value="$displayOa" >\n);
print qq(<input type="hidden" name="checkbox_cfp" value="$displayCfp">\n);
print qq(<input type="hidden" name="checkbox_afp" value="$displayAfp">\n);
print qq(<input type="hidden" name="checkbox_svm" value="$displaySvm">\n);
if ($displayOa) { &populateOaData(); }
if ($displayCfp) { &populateCfpData(); }
if ($displayAfp) { &populateAfpData(); }
if ($displaySvm) { &populateSvmData(); }
($oop, my $showJournal) = &getHtmlVar($query, "checkbox_journal"); unless ($showJournal) { $showJournal = ''; }
($oop, my $showPmid) = &getHtmlVar($query, "checkbox_pmid"); unless ($showPmid) { $showPmid = ''; }
($oop, my $showPdf) = &getHtmlVar($query, "checkbox_pdf"); unless ($showPdf) { $showPdf = ''; }
print qq(<input type="hidden" name="checkbox_journal" value="$showJournal">\n);
print qq(<input type="hidden" name="checkbox_pmid" value="$showPmid">\n);
print qq(<input type="hidden" name="checkbox_pdf" value="$showPdf">\n);
($oop, my $papersPerPage) = &getHtmlVar($query, "papers_per_page");
($oop, my $pageSelected) = &getHtmlVar($query, "select_page");
unless ($papersPerPage) { $papersPerPage = 10; }
unless ($pageSelected) { $pageSelected = 0; }
print qq(<input type="hidden" name="papers_per_page" value="$papersPerPage">\n);
my @headerRow = qw( paperID );
if ($showJournal) { push @headerRow, "journal"; &populateJournal(); }
if ($showPmid) { push @headerRow, "pmid"; &populatePmid(); }
if ($showPdf) { push @headerRow, "pdf"; &populatePdf(); }
The code now generates a hash of rows of results (not necessarily in the order submitted). Any paper that has data for the relevant datatype and flagging method, this data is then loaded into the appropriate column of each row.
my %trs; # td data for each table row
my %paperPosNegOkay; # papers that have positive-negative data okay, so show all svm results for that paper even if a given row isn't positive-negative okay
my %paperInfo; # for a joinkey, all the paper information about it to show in a big rowspan for that table row
my %allPaperData; # hash of datatype - joinkey for all posible queried data structures, to key off from this when there are no svm results for a data structure with data.
foreach my $datatype (keys %svmData) { foreach my $joinkey (keys %{ $svmData{$datatype} }) { $allPaperData{$joinkey}{$datatype}++; } }
foreach my $datatype (keys %curData) { foreach my $joinkey (keys %{ $curData{$datatype} }) { $allPaperData{$joinkey}{$datatype}++; } }
foreach my $datatype (keys %oaData) { foreach my $joinkey (keys %{ $oaData{$datatype} }) { $allPaperData{$joinkey}{$datatype}++; } }
foreach my $datatype (keys %cfpData) { foreach my $joinkey (keys %{ $cfpData{$datatype} }) { $allPaperData{$joinkey}{$datatype}++; } }
foreach my $datatype (keys %afpData) { foreach my $joinkey (keys %{ $afpData{$datatype} }) { $allPaperData{$joinkey}{$datatype}++; } }
my $trCounter = 0;
foreach my $joinkey (sort keys %curatablePapers) { # TODO curatablePapers or allPaperData that have some flag ?
next unless ($chosenPapers{$joinkey} || $chosenPapers{all});
push @{ $paperInfo{$joinkey} }, $joinkey;
my $journal = ''; my $pmid = ''; my $pdf = ''; my $primaryData = '';
if ($showJournal) {
if ($journal{$joinkey}) { $journal = $journal{$joinkey}; }
push @{ $paperInfo{$joinkey} }, $journal; }
if ($showPmid) {
if ($pmid{$joinkey}) { $pmid = $pmid{$joinkey}; }
push @{ $paperInfo{$joinkey} }, $pmid; }
if ($showPdf) {
if ($pdf{$joinkey}) { $pdf = $pdf{$joinkey}; }
push @{ $paperInfo{$joinkey} }, $pdf; }
foreach my $datatype (sort keys %{ $allPaperData{$joinkey} }) {
next unless ($chosenDatatypes{$datatype}); # show only results for selected datatype
my @dataRow = ( "$datatype" );
$trCounter++;
if ($displaySvm) {
my $svmResult = '';
if ($svmData{$datatype}{$joinkey}) { $svmResult = $svmData{$datatype}{$joinkey}; }
my $bgcolor = 'white';
if ($svmResult eq 'high') { $bgcolor = '#FFA0A0'; }
elsif ($svmResult eq 'medium') { $bgcolor = '#FFC8C8'; }
elsif ($svmResult eq 'low') { $bgcolor = '#FFE0E0'; }
$svmResult = qq(<span style="background-color: $bgcolor">$svmResult</span>);
push @dataRow, $svmResult;
} # if ($displaySvm)
if ($displayCfp) {
my $cfpResult = '';
if ($cfpData{$datatype}{$joinkey}) { $cfpResult = $cfpData{$datatype}{$joinkey}; }
push @dataRow, $cfpResult;
}
if ($displayAfp) {
my $afpResult = '';
if ($afpData{$datatype}{$joinkey}) { $afpResult = $afpData{$datatype}{$joinkey}; }
push @dataRow, $afpResult;
}
if ($displayOa) {
my $oaResult = 'oa_blank';
if ($oaData{$datatype}{$joinkey}) { $oaResult = $oaData{$datatype}{$joinkey}; }
push @dataRow, $oaResult;
}
my $thisCurator = ''; # curator in cur_curdata for this paper-datatype if it has a value
if ( $curData{$datatype}{$joinkey}{curator} ) { $thisCurator = $curData{$datatype}{$joinkey}{curator}; }
my $curatorSelectCurator = qq(<select name="select_curator_curator_$trCounter" size="1">\n<option value=""></option>\n);
foreach my $curator_two (keys %curators) { # display curators in alphabetical (tied hash) order, if IP matches existing ip record, select it
if ($thisCurator eq $curator_two) { $curatorSelectCurator .= qq(<option value="$curator_two" selected="selected">$curators{$curator_two}</option>\n); }
else { $curatorSelectCurator .= qq(<option value="$curator_two">$curators{$curator_two}</option>\n); } }
$curatorSelectCurator .= qq(</select>);
$curatorSelectCurator .= qq(<input type="hidden" name="joinkey_$trCounter" value="$joinkey" >); # these are required, arbitrarily added here
$curatorSelectCurator .= qq(<input type="hidden" name="datatype_$trCounter" value="$datatype">); # these are required, arbitrarily added here
push @dataRow, $curatorSelectCurator;
my $thisDonPosNeg = ''; if ( $curData{$datatype}{$joinkey}{donposneg} ) { $thisDonPosNeg = $curData{$datatype}{$joinkey}{donposneg}; }
my $curatorSelectDonposneg = qq(<select name="select_curator_donposneg_$trCounter">);
foreach my $donposneg (keys %donPosNegOptions) { # display curators in alphabetical (tied hash) order, if IP matches existing ip record, select it
if ($thisDonPosNeg eq $donposneg) { $curatorSelectDonposneg .= qq(<option value="$donposneg" selected="selected">$donPosNegOptions{$donposneg}</option>\n); }
else { $curatorSelectDonposneg .= qq(<option value="$donposneg" >$donPosNegOptions{$donposneg}</option>\n); } }
$curatorSelectDonposneg .= qq(</select>);
push @dataRow, $curatorSelectDonposneg;
my $thisSelComment = ''; if ( $curData{$datatype}{$joinkey}{selcomment} ) { $thisSelComment = $curData{$datatype}{$joinkey}{selcomment}; }
my $curatorSelectComment = qq(<select name="select_curator_comment_$trCounter">);
$curatorSelectComment .= qq(<option value="" ></option>\n);
foreach my $comment (keys %premadeComments) {
if ($thisSelComment eq $comment) { $curatorSelectComment .= qq(<option value="$comment" selected="selected">$premadeComments{$comment}</option>\n); }
else { $curatorSelectComment .= qq(<option value="$comment" >$premadeComments{$comment}</option>\n); } }
$curatorSelectComment .= qq(</select>);
push @dataRow, $curatorSelectComment;
my $txtcomment = ''; if ( $curData{$datatype}{$joinkey}{txtcomment} ) { $txtcomment = $curData{$datatype}{$joinkey}{txtcomment}; }
my $shortTxtComment = $txtcomment; unless ($shortTxtComment) { $shortTxtComment = ' '; }
if ($txtcomment =~ m/^(.{20})/) { $shortTxtComment = $1; $shortTxtComment .= '...'; }
my $curatorTextareaComment = qq(<div id="div_curator_comment_$trCounter" onclick="document.getElementById('div_curator_comment_$trCounter').style.display = 'none'; document.getElementById('textarea_curator_comment_$trCounter').style.display = ''; document.getElementById('textarea_curator_comment_$trCounter').focus();" >$shortTxtComment</div>\n);
$curatorTextareaComment .= qq(<textarea rows="4" cols="80" id="textarea_curator_comment_$trCounter" name="textarea_curator_comment_$trCounter" style="display:none" onblur="document.getElementById('div_curator_comment_$trCounter').style.display = ''; document.getElementById('textarea_curator_comment_$trCounter').style.display = 'none'; var divValue = document.getElementById('textarea_curator_comment_$trCounter').value; if (divValue === '') { divValue = ' '; } document.getElementById('div_curator_comment_$trCounter').innerHTML = divValue; ">$txtcomment</textarea>\n);
# $curatorTextareaComment .= qq(<textarea rows="4" cols="80" id="textarea_curator_comment_$trCounter" name="textarea_curator_comment_$trCounter" style="display:none" onblur="document.getElementById('div_curator_comment_$trCounter').style.display = ''; document.getElementById('textarea_curator_comment_$trCounter').style.display = 'none'; document.getElementById('div_curator_comment_$trCounter').innerHTML = document.getElementById('textarea_curator_comment_$trCounter').value.substring(0,20)">$txtcomment</textarea>\n); # to get the first 20 characters without adding ...
push @dataRow, $curatorTextareaComment;
$paperPosNegOkay{$joinkey}++; # all papers always okay for pos/neg since we no longer have pos/neg filtering 2012 11 08
my $trData = join"</td>$tdDot", @dataRow;
push @{ $trs{$joinkey} }, qq(${tdDot}$trData</td></tr>\n);
} # foreach my $datatype (sort keys %{ $allPaperData{$joinkey} })
} # foreach my $joinkey (sort keys %allPaperData)
print qq(<input type="hidden" name="trCounter" value="$trCounter">);
my $joinkeysAmount = scalar(keys %paperPosNegOkay);
my $pagesAmount = ceil($joinkeysAmount / $papersPerPage);
print qq(Page number <select name="select_page">);
for my $i (1 .. $pagesAmount) {
if ($i == $pageSelected) { print qq(<option selected="selected">$i</option>\n); }
else { print qq(<option>$i</option>\n); }
} # for my $i (1 .. $pagesAmount)
print qq(</select>);
print qq(<input type="submit" name="action" value="Get Results">\n);
print qq(amount of papers $joinkeysAmount<br/>\n);
print qq(<br />\n);
print qq(<table border="1">\n);
push @headerRow, "datatype";
if ($displaySvm) { push @headerRow, "SVM Prediction"; }
if ($displayCfp) { push @headerRow, "cfp value"; }
if ($displayAfp) { push @headerRow, "afp value"; }
if ($displayOa) { push @headerRow, "oa value"; }
push @headerRow, "curator"; push @headerRow, "new result"; push @headerRow, "select comment"; push @headerRow, "textarea comment";
my $headerRow = join"</th>$thDot", @headerRow;
$headerRow = qq(<tr>$thDot) . $headerRow . qq(</th></tr>);
print qq($headerRow\n);
my $papCount = 0;
my $papCountToSkip = 0; my $papToSkip = ($pageSelected - 1 ) * $papersPerPage;
foreach my $joinkey (sort keys %paperPosNegOkay) { # from all papers that have good positve-negative values, show all TRs
$papCountToSkip++; next if ($papCountToSkip <= $papToSkip); # skip entries until at the proper page
$papCount++;
last if ($papCount > $papersPerPage);
my $trsInPaperAmount = scalar @{ $trs{$joinkey} }; # amount of rows for a joinkey, make that the rowspan
my $firstTr = shift @{ $trs{$joinkey} }; # the first table row needs the paper info and rowspan
my $tdMultiRow = $tdDot; $tdMultiRow =~ s/>$/ rowspan="$trsInPaperAmount">/; # add the rowspan to the td style
my $paperInfoTds = join"</td>$tdMultiRow", @{ $paperInfo{$joinkey} }; # make paper info tds from %paperInfo
print qq(<tr>${tdMultiRow}$paperInfoTds</td>$firstTr\n); # print the first row which has paper info
foreach my $tr (@{ $trs{$joinkey} }) { print qq(<tr>$tr\n); } } # print other table rows without paper info
print qq(</table>\n);
print qq(<input type="submit" name="action" value="Submit New Results"><br/>\n);
&printFormClose();
} # sub getResults
Precanned Comments
In the Detailed Results of Papers page, curators have the option to select a comment from a drop down list of comments to apply to this paper in the context of the relevant datatype.
In the code, the comments are stored in a hash table called %premadeComments. The keys (stored in postgres) of these comments are only numbers, so the descriptions/titles can change or be updated and still apply retroactively.
Code:
sub populatePremadeComments {
$premadeComments{"1"} = "SVM Positive, Curation Negative";
$premadeComments{"2"} = "pre-made comment #2";
$premadeComments{"3"} = "pre-made comment #3";}
So, as of now:
| Key | Comment | | 1 | "SVM Positive, Curation Negative" | | 2 | "pre-made comment #2" | | 3 | "pre-made comment #3" |
Hence, if a completely new comment is desired, a new key will need to be made and there after associated with that new comment. Also, old keys should never be recycled and documentation describing what each key refers to should be maintained in this Wiki.
New Result
Each paper-datatype pair can be assigned a "New Result" indicating its status as curated (or not) or validated (or not), and if validated, positive or negative for the particular paper-datatype pair. These results can be entered via the Add Results Page or directly in the Detailed Results of Papers page via the "New Results" column. The code is below:
Code:
sub populateDonPosNegOptions {
$donPosNegOptions{""} = "";
$donPosNegOptions{"curated"} = "curated and positive";
$donPosNegOptions{"positive"} = "validated positive";
$donPosNegOptions{"negative"} = "validated negative";
$donPosNegOptions{"notvalidated"} = "not validated";}
where "curated", "positive", "negative", and "notvalidated" are the keys (for the %donPosNegOptions hash table in the form code) that will be stored in postgres and the corresponding values (e.g. "curated and positive") are what will be displayed on the form.
Note that "" and "not validated" represent no data for that paper-datatype pair, but "not validated" is present as an option to overwrite accidental validations (it is impossible to go back to a blank "" field via the form).
Datatypes
The form determines which datatypes exist via a 'populateDatatypes' subroutine in the form code. As of 12-5-2012, the form first collects all datatypes used in SVM from the 'cur_svmdata' postgres table (which, as of 12-5-2012, all also are identically named in the Author First Pass (AFP) and Curator First Pass (CFP) tables) and then supplements with other datatypes not in SVM but in AFP and CFP (as of 12-5-2012, all anatomy curation related datatypes) plus one additional datatype ("geneticablation") not in SVM, AFP, or CFP.
Here is the code:
sub populateDatatypes {
$result = $dbh->prepare( "SELECT DISTINCT(cur_datatype) FROM cur_svmdata " );
$result->execute() or die "Cannot prepare statement: $DBI::errstr\n";
while (my @row = $result->fetchrow) { $datatypesAfpCfp{$row[0]} = $row[0]; }
$datatypesAfpCfp{'blastomere'} = 'cellfunc';
$datatypesAfpCfp{'exprmosaic'} = 'siteaction';
$datatypesAfpCfp{'geneticmosaic'} = 'mosaic';
$datatypesAfpCfp{'laserablation'} = 'ablationdata';
foreach my $datatype (keys %datatypesAfpCfp) { $datatypes{$datatype}++; }
$datatypes{'geneticablation'}++;
} # sub populateDatatypes
As for the datatypes currently (12-5-2012) NOT in SVM but IN AFP and CFP, the datatype name is different between the Curation Status form and the AFP and CFP forms. So, the datatypes named "cellfunc", "siteaction", "mosaic", and "ablationdata" in the AFP and CFP tables are respectively named "blastomere", "exprmosaic", "geneticmosaic", "laserablation" in the Curation Status form.
The IMPORTANT thing here is: if, at some point, the datatypes are changed (added, renamed, etc.), and the code is not updated in kind, the form will likely break. Curators should tell Juancarlos/Chris/Daniela to update the code.
new datatypes should be accounted in this code :
- - no svm, no afp/cfp : add to %datatypes hash like 'geneticablation'.
- - no svm, yes afp/cfp : add to %datatypesAfpCfp + %datatypes hashes like 'blastomere'
- - yes svm, yes afp/cfp : add to code to populate cur_svmdata, which will populate in the SELECT query
- - yes svm, no afp/cfp : add to code to populate cur_svmdata, which will populate in the SELECT query, but also subsequently delete from %datatypesAfpCfp (to prevent a postgres query to a non-existing table which will crash the form)
Creating PDF links to papers
In the Detailed Results of Papers page, each paper ID is linked to its corresponding PDF document using the code below:
Code:
sub populatePdf {
$result = $dbh->prepare( "SELECT * FROM pap_electronic_path WHERE pap_electronic_path IS NOT NULL");
$result->execute() or die "Cannot prepare statement: $DBI::errstr\n";
my %temp;
while (my @row = $result->fetchrow) {
my ($data, $isPdf) = &makePdfLinkFromPath($row[1]);
$temp{$row[0]}{$isPdf}{$data}++; }
foreach my $joinkey (sort keys %temp) {
my @pdfs;
foreach my $isPdf (reverse sort keys %{ $temp{$joinkey} }) {
foreach my $pdfLink (sort keys %{ $temp{$joinkey}{$isPdf} }) {
push @pdfs, $pdfLink; } }
my ($pdfs) = join"<br/>", @pdfs;
$pdf{$joinkey} = $pdfs;
} # foreach my $joinkey (sort keys %temp)
} # sub populatePdf
sub makePdfLinkFromPath {
my ($path) = shift;
my ($pdf) = $path =~ m/\/([^\/]*)$/;
my $isPdf = 0; if ($pdf =~ m/\.pdf$/) { $isPdf++; } # kimberly wants .pdf files on top, so need to flag to sort
my $link = 'http://tazendra.caltech.edu/~acedb/daniel/' . $pdf;
my $data = "<a href=\"$link\" target=\"new\">$pdf</a>"; return ($data, $isPdf); }
Note the table name ("pap_electronic_path"), the URL path ("http://tazendra.caltech.edu/~acedb/daniel/"), and (because of the code 'target=\"new\"') that the link will open a new window or tab. Also note that opening another link on the original page (e.g. Detailed Results of Papers page) will open that link in that same new window/tab, clearing out what you had opened previously.
Creating hyperlinks to PubMed paper pages
In the Detailed Results of Papers page each PubMed ID is linked to its corresponding PubMed webpage using the code below:
Code:
sub populatePmid {
$result = $dbh->prepare( "SELECT * FROM pap_identifier WHERE pap_identifier ~ 'pmid'" );
$result->execute() or die "Cannot prepare statement: $DBI::errstr\n";
my %temp;
while (my @row = $result->fetchrow) { if ($row[0]) {
my ($data) = &makeNcbiLinkFromPmid($row[1]);
$temp{$row[0]}{$data}++; } }
foreach my $joinkey (sort keys %temp) {
my ($pmids) = join"<br/>", keys %{ $temp{$joinkey} };
$pmid{$joinkey} = $pmids;
} # foreach my $joinkey (sort keys %temp)
} # sub populatePmid
sub makeNcbiLinkFromPmid {
my $pmid = shift;
my ($id) = $pmid =~ m/(\d+)/;
my $link = 'http://www.ncbi.nlm.nih.gov/pubmed/' . $id;
my $data = "<a href=\"$link\" target=\"new\">$pmid</a>"; return $data; }
Note the table name ("pap_identifier"), the table specifier ("WHERE pap_identifier ~ 'pmid'"), the URL path ("http://www.ncbi.nlm.nih.gov/pubmed/"), and (because of the code 'target=\"new\"') that the link will open a new window or tab. Also note that opening another link on the original page (e.g. Detailed Results of Papers page) will open that link in that same new window/tab, clearing out what you had opened previously.
Populating the Journal Names
Journal names for each paper are populated via the following code:
sub populateJournal {
$result = $dbh->prepare( "SELECT * FROM pap_journal WHERE pap_journal IS NOT NULL" );
$result->execute() or die "Cannot prepare statement: $DBI::errstr\n";
while (my @row = $result->fetchrow) { if ($row[0]) { $journal{$row[0]} = $row[1]; } }
} # sub populateJournal
Note the table "pap_journal".
Loading Data into the Form
On the Curation Statistics Options Page, the Specific Paper Page, or the Prepopulated Specific Papers Page, curators have the option to specify what flagging methods (SVM, AFP, and/or CFP), curation sources (Ontology Annotator or cur_curdata [which is the data generated from this form]), and/or datatypes (e.g. geneint, rnai) they would like to view.
There are separate hashes for storing the different types of data, all of which have a key of datatype, subkey paperID, sub-subkeys of other things depending on the hash (see individual subsections below).
There is an option to select specific datatype, in which case only the data for those datatypes is loaded. Similarly if only some paperIDs have been selected, only those paperIDs are loaded.
Loading curatable papers
Only papers that have a 'valid' pap_status value and a 'primary' pap_primary_data value are considered curatable. These are stored in the %curatablePapers hash. ( paperID => status )
sub populateCuratablePapers {
my $query = "SELECT * FROM pap_status WHERE pap_status = 'valid' AND joinkey IN (SELECT joinkey FROM pap_primary_data WHERE pap_primary_data = 'primary')";
$result = $dbh->prepare( $query );
$result->execute() or die "Cannot prepare statement: $DBI::errstr\n";
while (my @row = $result->fetchrow) { $curatablePapers{$row[0]} = $row[1]; }
} # sub populateCuratablePapers
Loading afp_ data
Populate %afpEmailed, %afpData, %afpFlagged, %afpPos, %afpNeg.
for each of the chosen datatypes, if they are allowed in %datatypeAfpCfp, query the corresponding afp_ postgres table, and if it's a curatable paper store the value in the %afpData hash (datatype, paper ID => AFP result).
Query afp_email and if it's a curatable paper store in %afpEmailed hash ( paperID => 1 ) for afp emailed statistics.
Query afp_lasttouched to see if a paper has been flagged for afp. Skip if it's not a curatable paper. For all %chosenDatatypes store in %afpFlagged ( datatype, paperID => 1 )
For each of the %afpFlagged datatypes that have been chosen (%chosenDatatypes), if there is an %afpData value, store in %afpPos hash ( positive flag for afp ), otherwise store in %afpNeg hash (negative flag for afp ) ( datatype, paperID => 1 )
sub populateAfpData {
foreach my $datatype (sort keys %chosenDatatypes) {
next unless $datatypesAfpCfp{$datatype};
my $pgtable_datatype = $datatypesAfpCfp{$datatype};
$result = $dbh->prepare( "SELECT * FROM afp_$pgtable_datatype" );
$result->execute() or die "Cannot prepare statement: $DBI::errstr\n";
while (my @row = $result->fetchrow) {
next unless ($curatablePapers{$row[0]});
$afpData{$datatype}{$row[0]} = $row[1]; }
} # foreach my $datatype (sort keys %chosenDatatypes)
$result = $dbh->prepare( "SELECT * FROM afp_email" );
$result->execute() or die "Cannot prepare statement: $DBI::errstr\n";
while (my @row = $result->fetchrow) {
next unless ($curatablePapers{$row[0]});
$afpEmailed{$row[0]}++; }
$result = $dbh->prepare( "SELECT * FROM afp_lasttouched" );
$result->execute() or die "Cannot prepare statement: $DBI::errstr\n";
while (my @row = $result->fetchrow) {
next unless ($curatablePapers{$row[0]});
foreach my $datatype (sort keys %chosenDatatypes) {
$afpFlagged{$datatype}{$row[0]}++; } }
foreach my $datatype (sort keys %chosenDatatypes) {
foreach my $joinkey (sort keys %{ $afpFlagged{$datatype} }) {
if ($afpData{$datatype}{$joinkey}) { $afpPos{$datatype}{$joinkey}++; }
else { $afpNeg{$datatype}{$joinkey}++; } } }
} # sub populateAfpData
Loading cfp_ data
Populate %cfpData, %cfpFlagged, %cfpPos, %cfpNeg.
for each of the chosen datatypes, if they are allowed in %datatypeAfpCfp, query the corresponding cfp_ postgres table, and if it's a curatable paper store the value in the %cfpData hash (datatype, paper ID => CFP result).
Query cfp_curator to see if a paper has been flagged for cfp. Skip if it's not a curatable paper. For all %chosenDatatypes store in %cfpFlagged ( datatype, paperID => 1 )
For each of the %cfpFlagged datatypes that have been chosen (%chosenDatatypes), if there is an %cfpData value, store in %cfpPos hash ( positive flag for cfp ), otherwise store in %cfpNeg hash (negative flag for cfp ) ( datatype, paperID => 1 )
sub populateCfpData {
foreach my $datatype (sort keys %chosenDatatypes) {
next unless $datatypesAfpCfp{$datatype};
my $pgtable_datatype = $datatypesAfpCfp{$datatype};
$result = $dbh->prepare( "SELECT * FROM cfp_$pgtable_datatype" );
$result->execute() or die "Cannot prepare statement: $DBI::errstr\n";
while (my @row = $result->fetchrow) {
next unless ($curatablePapers{$row[0]});
$cfpData{$datatype}{$row[0]} = $row[1]; }
} # foreach my $datatype (sort keys %chosenDatatypes)
$result = $dbh->prepare( "SELECT * FROM cfp_curator" );
$result->execute() or die "Cannot prepare statement: $DBI::errstr\n";
while (my @row = $result->fetchrow) {
next unless ($curatablePapers{$row[0]});
foreach my $datatype (sort keys %chosenDatatypes) {
$cfpFlagged{$datatype}{$row[0]}++; } }
foreach my $datatype (sort keys %chosenDatatypes) {
foreach my $joinkey (sort keys %{ $cfpFlagged{$datatype} }) {
if ($cfpData{$datatype}{$joinkey}) { $cfpPos{$datatype}{$joinkey}++; }
else { $cfpNeg{$datatype}{$joinkey}++; } } }
} # sub populateCfpData
Loading svm data
Populate %svmData hash.
For each of the chosen datatypes, query the cur_svmdata table where cur_datatype is that datatype, and sort by cur_date so that we always have the latest value for a given paper-datatype pair. The svm result is the 4th column, the paper ID is the first column. skip papers that are not %curatablePapers. store in %svmData ( datatype, paper => svm_result ). cur_svmdata could have multiple results for a given paper-datatype pair, we'll consider only the most recent result (by the directory name/date on Yuling's machine).
sub populateSvmData {
# $result = $dbh->prepare( "SELECT * FROM cur_svmdata ORDER BY cur_datatype, cur_date" ); # always doing for all datatypes vs looping for chosen takes 4.66vs 2.74 secs
foreach my $datatype (sort keys %chosenDatatypes) {
$result = $dbh->prepare( "SELECT * FROM cur_svmdata WHERE cur_datatype = '$datatype' ORDER BY cur_date" );
# table stores multiple dates for same paper-datatype in case we want to see multiple results later. if it didn't and we didn't order it would take 2.05 vs 2.74 secs, so not worth changing the way we're storing data
$result->execute() or die "Cannot prepare statement: $DBI::errstr\n";
while (my @row = $result->fetchrow) {
my $joinkey = $row[0]; my $svmdata = $row[3];
next unless ($curatablePapers{$row[0]});
$svmData{$datatype}{$joinkey} = $svmdata; } }
} # sub populateSvmData
Loading OA data
Populate %objsCurated and %oaData hashes.
Each datatype is stored in different tables and has to be queried separately. The queries are mostly the same.
if ($chosenDatatypes{'newmutant'}) {
$result = $dbh->prepare( "SELECT * FROM app_variation" );
$result->execute() or die "Cannot prepare statement: $DBI::errstr\n";
while (my @row = $result->fetchrow) { $objsCurated{'newmutant'}{$row[1]}++; }
$result = $dbh->prepare( "SELECT * FROM app_paper" );
$result->execute() or die "Cannot prepare statement: $DBI::errstr\n";
while (my @row = $result->fetchrow) {
my (@papers) = $row[1] =~ m/WBPaper(\d+)/g;
foreach my $paper (@papers) {
$oaData{'newmutant'}{$paper} = 'curated'; } } }
and similarly for other datatypes.
The example above is for the datatype 'newmutant'. If that datatype is a %chosenDatatypes, query app_variation and store in %objsCurated ( datatype, object => 1 ), then query app_paper matching for WBPaper IDs, and associating to %oaData ( datatype, paperID => 'curated' ).
For other datatypes :
- overexpr : objects from app_transgene ; %oaData from app_paper WHERE joinkey IN (SELECT joinkey FROM app_transgene WHERE app_transgene IS NOT NULL AND app_transgene != ), meaning papers where the postgresID has a corresponding transgene that exists in app_transgene.
- antibody : objects from abp_name ; %oaData from abp_paper
- otherexpr : objects from exp_name ; %oaData from exp_paper
- genereg : objects from grg_name; %oaData from grg_paper
- geneint : objects from int_name; %oaData from int_paper
- rnai : objects from rna_name; %oaData from rna_paper
- blastomere : objects from wbb_wbbtf WHERE joinkey IN (SELECT joinkey FROM wbb_assay WHERE wbb_assay = 'Blastomere_isolation') ; %oaData from wbb_reference WHERE joinkey IN (SELECT joinkey FROM wbb_assay WHERE wbb_assay = 'Blastomere_isolation')
- exprmosaic : objects from wbb_wbbtf WHERE joinkey IN (SELECT joinkey FROM wbb_assay WHERE wbb_assay = 'Expression_mosaic') ; %oaData from wbb_reference WHERE joinkey IN (SELECT joinkey FROM wbb_assay WHERE wbb_assay = 'Expression_mosaic')
- geneticablation : objects from wbb_wbbtf WHERE joinkey IN (SELECT joinkey FROM wbb_assay WHERE wbb_assay = 'Genetic_ablation') ; %oaData from wbb_reference WHERE joinkey IN (SELECT joinkey FROM wbb_assay WHERE wbb_assay = 'Genetic_ablation')
- geneticmosaic : objects from wbb_wbbtf WHERE joinkey IN (SELECT joinkey FROM wbb_assay WHERE wbb_assay = 'Genetic_mosaic') ; %oaData from wbb_reference WHERE joinkey IN (SELECT joinkey FROM wbb_assay WHERE wbb_assay = 'Genetic_mosaic')
- laserablation : objects from wbb_wbbtf WHERE joinkey IN (SELECT joinkey FROM wbb_assay WHERE wbb_assay = 'Laser_ablation') ; %oaData from wbb_reference WHERE joinkey IN (SELECT joinkey FROM wbb_assay WHERE wbb_assay = 'Laser_ablation')
Loading cur_curdata
cur_curdata: this captures all data entered through this form, meaning paper ID, datatype, curator ID, validation status (e.g. "curated and positive"), pre-canned comment, and/or free text comment (and timestamp). Note: this table only stores data (and associated paper-datatype pairs) that has been manually entered through this form.
Code:
sub populateCurCurData {
$result = $dbh->prepare( "SELECT * FROM cur_curdata ORDER BY cur_timestamp" ); # in case multiple values get in for a paper-datatype (shouldn't happen), keep the latest
$result->execute() or die "Cannot prepare statement: $DBI::errstr\n";
while (my @row = $result->fetchrow) {
next unless ($chosenPapers{$row[0]} || $chosenPapers{all});
next unless ($chosenDatatypes{$row[1]});
$curData{$row[1]}{$row[0]}{curator} = $row[2];
$curData{$row[1]}{$row[0]}{donposneg} = $row[3];
$curData{$row[1]}{$row[0]}{selcomment} = $row[4];
$curData{$row[1]}{$row[0]}{txtcomment} = $row[5];
$curData{$row[1]}{$row[0]}{timestamp} = $row[6]; }
} # sub populateCurCurData
When populating curator data from curation status, read the cur_curdata postgres table, skip datatypes that were not chosen, skip papers that were not chosen.
Store data in the %curData hash, key is datatype, subkey is paperID, then valuekeys are curator, donposneg (curator result of curated, validatedPos, validatedNeg, notValidated), select comment, text comment, timestamp.
cur_curdata can only have one result for a specific paper-datatype pair, if a new result is entered it will overwrite the previous result.
Processing curated data
The following subroutine will process cur_curdata and oaData into %valCur %valPos %valNeg and into %conflict which has the paper-datatypes that have multiple values, which correspond to a datatype-paper pair's validated+curated, validated+positive, validated+negative.
If a paper has been curated for a datatype, the paper enters into the %valCur AND the %valPos hashes; if it has been validated positive but NOT curated it goes into %valPos ONLY; and if it has been validated negative it will go into %valNeg.
sub populateCuratedPapers {
my ($showTimes, $start, $end, $diff) = (0, '', '', '');
if ($showTimes) { $start = time; }
&populateCurCurData();
if ($showTimes) { $end = time; $diff = $end - $start; $start = time; print "IN populateCuratedPapers populateCurCurData $diff<br>"; }
&populateOa(); # $oaData{datatype}{joinkey} = 'positive';
if ($showTimes) { $end = time; $diff = $end - $start; $start = time; print "IN populateCuratedPapers populateOa $diff<br>"; }
my %allCuratorValues; # $allCuratorValues{datatype}{joinkey} = 0 | 1+
foreach my $datatype (sort keys %oaData) {
foreach my $joinkey (sort keys %{ $oaData{$datatype} }) {
$allCuratorValues{$joinkey}{$datatype}{curated}++; } } # validated positive and curated
foreach my $datatype (sort keys %curData) {
foreach my $joinkey (sort keys %{ $curData{$datatype} }) {
$allCuratorValues{$joinkey}{$datatype}{ $curData{$datatype}{$joinkey}{donposneg} }++; } }
foreach my $joinkey (sort keys %allCuratorValues) {
foreach my $datatype (sort keys %{ $allCuratorValues{$joinkey} }) {
my @values = keys %{ $allCuratorValues{$joinkey}{$datatype} };
if (scalar @values > 1) { $conflict{$datatype}{$joinkey}++; }
else {
my $value = shift @values;
$validated{$datatype}{$joinkey} = $value;
if ($value eq 'curated') { $valPos{$datatype}{$joinkey} = $value; $valCur{$datatype}{$joinkey} = $value; }
elsif ($value eq 'positive') { $valPos{$datatype}{$joinkey} = $value; }
elsif ($value eq 'negative') { $valNeg{$datatype}{$joinkey} = $value; }
} } }
if ($showTimes) { $end = time; $diff = $end - $start; $start = time; print "IN populateCuratedPapers categorizing hash $diff<br>"; }
} # sub populateCuratedPapers
Curation Statistics Calculations
The way that each value is calculated for Curation Statistics table is based on what papers (or, more specifically, paper IDs) populate each of a number of tables. The following hash tables capture validation status:
%valCur - All papers that have been curated for a given datatype %valPos - All papers that have been validated positive for a given datatype, but not yet curated %valNeg - All papers that have been validated negative for a given datatype
When determining, for a particular flagging method, the validation and curation statistics with respect to flagging status, these tables are compared to the table for flagging results to generate the numbers for the Curation Statistics table. So, for AFP Positives for example, the following logic is performed to determine the indicated values (list of papers), per datatype:
AFP positive (%afpPos) AFP positive validated (%afpPosVal) : %afpPos AND (%valNeg OR %valPos) AFP positive validated false positive (%afpPosFP) : %afpPos AND %valNeg AFP positive validated true positive (%afpPosTP) : %afpPos AND %valPos AFP positive validated true positive curated (%afpPosTpCur) : %afpPos AND %valPos AND %valCur <Note: the %valPOS is redundant> AFP positive validated true positive not curated (%afpPosTpNC) : %afpPos AND (%valPos NOT %valCur) AFP positive not validated (%afpPosNV) : %afpPos NOT (%valNeg OR %valPos) AFP positive not curated (%afpPosNC) : (%afpPos AND (%valPos NOT %valCur)) OR (%afpPos NOT (%valNeg OR %valPos))
which are determined by the following section of code:
sub getCurationStatisticsAfpPos {
my ($datatypesToShow_ref) = @_;
my @datatypesToShow = @$datatypesToShow_ref;
my %afpPosNV; my %afpPosVal; my %afpPosFP; my %afpPosTP; my %afpPosTpCur; my %afpPosTpNC; my %afpPosNC;
# positive and : not validated, validated, false positive, true positive, TP curated, TP not curated, not curated minus validated negative OR not validated + TP not curated
foreach my $datatype (@datatypesToShow) {
foreach my $joinkey (sort keys %{ $afpPos{$datatype} }) {
if ($valPos{$datatype}{$joinkey}) { $afpPosTP{$datatype}{$joinkey}++; $afpPosVal{$datatype}{$joinkey}++;
if ($valCur{$datatype}{$joinkey}) { $afpPosTpCur{$datatype}{$joinkey}++; }
else { $afpPosTpNC{$datatype}{$joinkey}++; $afpPosNC{$datatype}{$joinkey}++; } }
elsif ($valNeg{$datatype}{$joinkey}) { $afpPosFP{$datatype}{$joinkey}++; $afpPosVal{$datatype}{$joinkey}++; }
else { $afpPosNV{$datatype}{$joinkey}++; $afpPosNC{$datatype}{$joinkey}++; } } }
tie %{ $curStats{'afp'}{'pos'} }, "Tie::IxHash";
foreach my $datatype (@datatypesToShow) {
my $countAfpFlagged = scalar keys %{ $afpFlagged{$datatype} };
my $countAfpPos = scalar keys %{ $afpPos{$datatype} };
my $ratio = 0;
if ($countAfpFlagged > 0) { $ratio = $countAfpPos / $countAfpFlagged * 100; $ratio = FormatSigFigs($ratio, 2); }
foreach my $joinkey (keys %{ $afpPos{$datatype} }) { $curStats{'afp'}{'pos'}{$datatype}{papers}{$joinkey}++;
$curStats{'any'}{'pos'}{$datatype}{papers}{$joinkey}++; }
$curStats{'afp'}{'pos'}{$datatype}{'countPap'} = scalar keys %{ $afpPos{$datatype} };
$curStats{'afp'}{'pos'}{$datatype}{'ratio'} = $ratio;
my $countAfpPosVal = scalar keys %{ $afpPosVal{$datatype} };
$ratio = 0;
if ($countAfpPos > 0) { $ratio = $countAfpPosVal / $countAfpPos * 100; $ratio = FormatSigFigs($ratio, 2); }
foreach my $joinkey (keys %{ $afpPosVal{$datatype} }) { $curStats{'afp'}{'pos'}{'val'}{$datatype}{papers}{$joinkey}++;
$curStats{'any'}{'pos'}{'val'}{$datatype}{papers}{$joinkey}++; }
$curStats{'afp'}{'pos'}{'val'}{$datatype}{'countPap'} = scalar keys %{ $afpPosVal{$datatype} };
$curStats{'afp'}{'pos'}{'val'}{$datatype}{'ratio'} = $ratio;
my $countAfpPosTP = scalar keys %{ $afpPosTP{$datatype} };
$ratio = 0;
if ($countAfpPosVal > 0) { $ratio = $countAfpPosTP / $countAfpPosVal * 100; $ratio = FormatSigFigs($ratio, 2); }
foreach my $joinkey (keys %{ $afpPosTP{$datatype} }) { $curStats{'afp'}{'pos'}{'val'}{'tp'}{$datatype}{papers}{$joinkey}++;
$curStats{'any'}{'pos'}{'val'}{'tp'}{$datatype}{papers}{$joinkey}++; }
$curStats{'afp'}{'pos'}{'val'}{'tp'}{$datatype}{'countPap'} = scalar keys %{ $afpPosTP{$datatype} };
$curStats{'afp'}{'pos'}{'val'}{'tp'}{$datatype}{'ratio'} = $ratio;
my $countAfpPosTpCur = scalar keys %{ $afpPosTpCur{$datatype} };
$ratio = 0;
if ($countAfpPosTP > 0) { $ratio = $countAfpPosTpCur / $countAfpPosTP * 100; $ratio = FormatSigFigs($ratio, 2); }
foreach my $joinkey (keys %{ $afpPosTpCur{$datatype} }) { $curStats{'afp'}{'pos'}{'val'}{'tp'}{'cur'}{$datatype}{papers}{$joinkey}++;
$curStats{'any'}{'pos'}{'val'}{'tp'}{'cur'}{$datatype}{papers}{$joinkey}++; }
$curStats{'afp'}{'pos'}{'val'}{'tp'}{'cur'}{$datatype}{'countPap'} = scalar keys %{ $afpPosTpCur{$datatype} };
$curStats{'afp'}{'pos'}{'val'}{'tp'}{'cur'}{$datatype}{'ratio'} = $ratio;
my $countAfpPosTpNC = scalar keys %{ $afpPosTpNC{$datatype} };
$ratio = 0;
if ($countAfpPosTP > 0) { $ratio = $countAfpPosTpNC / $countAfpPosTP * 100; $ratio = FormatSigFigs($ratio, 2); }
foreach my $joinkey (keys %{ $afpPosTpNC{$datatype} }) { $curStats{'afp'}{'pos'}{'val'}{'tp'}{'ncur'}{$datatype}{papers}{$joinkey}++;
$curStats{'any'}{'pos'}{'val'}{'tp'}{'ncur'}{$datatype}{papers}{$joinkey}++; }
$curStats{'afp'}{'pos'}{'val'}{'tp'}{'ncur'}{$datatype}{'countPap'} = scalar keys %{ $afpPosTpNC{$datatype} };
$curStats{'afp'}{'pos'}{'val'}{'tp'}{'ncur'}{$datatype}{'ratio'} = $ratio;
my $countAfpPosFP = scalar keys %{ $afpPosFP{$datatype} };
$ratio = 0;
if ($countAfpPosVal > 0) { $ratio = $countAfpPosFP / $countAfpPosVal * 100; $ratio = FormatSigFigs($ratio, 2); }
foreach my $joinkey (keys %{ $afpPosFP{$datatype} }) { $curStats{'afp'}{'pos'}{'val'}{'fp'}{$datatype}{papers}{$joinkey}++;
$curStats{'any'}{'pos'}{'val'}{'fp'}{$datatype}{papers}{$joinkey}++; }
$curStats{'afp'}{'pos'}{'val'}{'fp'}{$datatype}{'countPap'} = scalar keys %{ $afpPosFP{$datatype} };
$curStats{'afp'}{'pos'}{'val'}{'fp'}{$datatype}{'ratio'} = $ratio;
my $countAfpPosNV = scalar keys %{ $afpPosNV{$datatype} };
$ratio = 0;
if ($countAfpPos > 0) { $ratio = $countAfpPosNV / $countAfpPos * 100; $ratio = FormatSigFigs($ratio, 2); }
foreach my $joinkey (keys %{ $afpPosNV{$datatype} }) { $curStats{'afp'}{'pos'}{'nval'}{$datatype}{papers}{$joinkey}++;
$curStats{'any'}{'pos'}{'nval'}{$datatype}{papers}{$joinkey}++; }
$curStats{'afp'}{'pos'}{'nval'}{$datatype}{'countPap'} = scalar keys %{ $afpPosNV{$datatype} };
$curStats{'afp'}{'pos'}{'nval'}{$datatype}{'ratio'} = $ratio;
my $countAfpPosNC = scalar keys %{ $afpPosNC{$datatype} };
$ratio = 0;
if ($countAfpPos > 0) { $ratio = $countAfpPosNC / $countAfpPos * 100; $ratio = FormatSigFigs($ratio, 2); }
foreach my $joinkey (keys %{ $afpPosNC{$datatype} }) { $curStats{'afp'}{'pos'}{'ncur'}{$datatype}{papers}{$joinkey}++;
$curStats{'any'}{'pos'}{'ncur'}{$datatype}{papers}{$joinkey}++; }
$curStats{'afp'}{'pos'}{'ncur'}{$datatype}{'countPap'} = scalar keys %{ $afpPosNC{$datatype} };
$curStats{'afp'}{'pos'}{'ncur'}{$datatype}{'ratio'} = $ratio;
} # foreach my $datatype (@datatypesToShow)
} # sub getCurationStatisticsAfpPos
For AFP Negatives, the following logic is performed to determine the indicated values (list of papers), per datatype:
AFP negative (%afpNeg) AFP negative validated (%afpNegVal) : %afpNeg AND (%valNeg OR %valPos) AFP negative validated true negative (%afpNegTN) : %afpNeg AND %valNeg AFP negative validated false negative (%afpNegFN) : %afpNeg AND %valPos AFP negative validated false negative curated (%afpNegFnCur) : %afpNeg AND %valPos AND %valCur <Note: the %valPOS is redundant> AFP negative validated false negative not curated (%afpNegFnNC) : %afpNeg AND %valPos NOT %valCur AFP negative not validated (%afpNegNV) : %afpNeg NOT (%valNeg OR %valPos) AFP negative not curated (%afpNegNC) : (%afpNeg AND (%valPos NOT %valCur)) OR (%afpNeg NOT (%valPos OR %valNeg))
which are determined by the following section of code:
sub getCurationStatisticsAfpNeg {
my ($datatypesToShow_ref) = @_;
my @datatypesToShow = @$datatypesToShow_ref;
my %afpNegNV; my %afpNegVal; my %afpNegTN; my %afpNegFN; my %afpNegFnCur; my %afpNegFnNC; my %afpNegNC;
# negative and : not validated, validated, true negative, false negative, FN curated, FN not curated, not curated minus validated negative OR not validated + FN not curated
foreach my $datatype (@datatypesToShow) {
foreach my $joinkey (sort keys %{ $afpNeg{$datatype} }) {
if ($valNeg{$datatype}{$joinkey}) { $afpNegTN{$datatype}{$joinkey}++; $afpNegVal{$datatype}{$joinkey}++; }
elsif ($valPos{$datatype}{$joinkey}) { $afpNegFN{$datatype}{$joinkey}++; $afpNegVal{$datatype}{$joinkey}++;
if ($valCur{$datatype}{$joinkey}) { $afpNegFnCur{$datatype}{$joinkey}++; }
else { $afpNegFnNC{$datatype}{$joinkey}++; $afpNegNC{$datatype}{$joinkey}++; } }
else { $afpNegNV{$datatype}{$joinkey}++; $afpNegNC{$datatype}{$joinkey}++; } } }
tie %{ $curStats{'afp'}{'neg'} }, "Tie::IxHash";
foreach my $datatype (@datatypesToShow) {
my $countAfpFlagged = scalar keys %{ $afpFlagged{$datatype} };
my $countAfpNeg = scalar keys %{ $afpNeg{$datatype} };
my $ratio = 0;
if ($countAfpFlagged > 0) { $ratio = $countAfpNeg / $countAfpFlagged * 100; $ratio = FormatSigFigs($ratio, 2); } # ($ratio) = &roundToPlaces($ratio, 2); # $ratio = sprintf "%.2f", $ratio;
foreach my $joinkey (keys %{ $afpNeg{$datatype} }) { $curStats{'afp'}{'neg'}{$datatype}{papers}{$joinkey}++; }
$curStats{'afp'}{'neg'}{$datatype}{'countPap'} = scalar keys %{ $afpNeg{$datatype} };
$curStats{'afp'}{'neg'}{$datatype}{'ratio'} = $ratio;
my $countAfpNegVal = scalar keys %{ $afpNegVal{$datatype} };
$ratio = 0;
if ($countAfpNeg > 0) { $ratio = $countAfpNegVal / $countAfpNeg * 100; $ratio = FormatSigFigs($ratio, 2); }
foreach my $joinkey (keys %{ $afpNegVal{$datatype} }) { $curStats{'afp'}{'neg'}{'val'}{$datatype}{papers}{$joinkey}++; }
$curStats{'afp'}{'neg'}{'val'}{$datatype}{'countPap'} = scalar keys %{ $afpNegVal{$datatype} };
$curStats{'afp'}{'neg'}{'val'}{$datatype}{'ratio'} = $ratio;
my $countAfpNegTN = scalar keys %{ $afpNegTN{$datatype} };
$ratio = 0;
if ($countAfpNegVal > 0) { $ratio = $countAfpNegTN / $countAfpNegVal * 100; $ratio = FormatSigFigs($ratio, 2); }
foreach my $joinkey (keys %{ $afpNegTN{$datatype} }) { $curStats{'afp'}{'neg'}{'val'}{'tn'}{$datatype}{papers}{$joinkey}++; }
$curStats{'afp'}{'neg'}{'val'}{'tn'}{$datatype}{'countPap'} = scalar keys %{ $afpNegTN{$datatype} };
$curStats{'afp'}{'neg'}{'val'}{'tn'}{$datatype}{'ratio'} = $ratio;
my $countAfpNegFN = scalar keys %{ $afpNegFN{$datatype} };
$ratio = 0;
if ($countAfpNegVal > 0) { $ratio = $countAfpNegFN / $countAfpNegVal * 100; $ratio = FormatSigFigs($ratio, 2); }
foreach my $joinkey (keys %{ $afpNegFN{$datatype} }) { $curStats{'afp'}{'neg'}{'val'}{'fn'}{$datatype}{papers}{$joinkey}++; }
$curStats{'afp'}{'neg'}{'val'}{'fn'}{$datatype}{'countPap'} = scalar keys %{ $afpNegFN{$datatype} };
$curStats{'afp'}{'neg'}{'val'}{'fn'}{$datatype}{'ratio'} = $ratio;
my $countAfpNegFnCur = scalar keys %{ $afpNegFnCur{$datatype} };
$ratio = 0;
if ($countAfpNegFN > 0) { $ratio = $countAfpNegFnCur / $countAfpNegFN * 100; $ratio = FormatSigFigs($ratio, 2); }
foreach my $joinkey (keys %{ $afpNegFnCur{$datatype} }) { $curStats{'afp'}{'neg'}{'val'}{'fn'}{'cur'}{$datatype}{papers}{$joinkey}++; }
$curStats{'afp'}{'neg'}{'val'}{'fn'}{'cur'}{$datatype}{'countPap'} = scalar keys %{ $afpNegFnCur{$datatype} };
$curStats{'afp'}{'neg'}{'val'}{'fn'}{'cur'}{$datatype}{'ratio'} = $ratio;
my $countAfpNegFnNC = scalar keys %{ $afpNegFnNC{$datatype} };
$ratio = 0;
if ($countAfpNegFN > 0) { $ratio = $countAfpNegFnNC / $countAfpNegFN * 100; $ratio = FormatSigFigs($ratio, 2); }
foreach my $joinkey (keys %{ $afpNegFnNC{$datatype} }) { $curStats{'afp'}{'neg'}{'val'}{'fn'}{'ncur'}{$datatype}{papers}{$joinkey}++; }
$curStats{'afp'}{'neg'}{'val'}{'fn'}{'ncur'}{$datatype}{'countPap'} = scalar keys %{ $afpNegFnNC{$datatype} };
$curStats{'afp'}{'neg'}{'val'}{'fn'}{'ncur'}{$datatype}{'ratio'} = $ratio;
my $countAfpNegNV = scalar keys %{ $afpNegNV{$datatype} };
$ratio = 0;
if ($countAfpNeg > 0) { $ratio = $countAfpNegNV / $countAfpNeg * 100; $ratio = FormatSigFigs($ratio, 2); }
foreach my $joinkey (keys %{ $afpNegNV{$datatype} }) { $curStats{'afp'}{'neg'}{'nval'}{$datatype}{papers}{$joinkey}++; }
$curStats{'afp'}{'neg'}{'nval'}{$datatype}{'countPap'} = scalar keys %{ $afpNegNV{$datatype} };
$curStats{'afp'}{'neg'}{'nval'}{$datatype}{'ratio'} = $ratio;
my $countAfpNegNC = scalar keys %{ $afpNegNC{$datatype} };
$ratio = 0;
if ($countAfpNeg > 0) { $ratio = $countAfpNegNC / $countAfpNeg * 100; $ratio = FormatSigFigs($ratio, 2); }
foreach my $joinkey (keys %{ $afpNegNC{$datatype} }) { $curStats{'afp'}{'neg'}{'ncur'}{$datatype}{papers}{$joinkey}++; }
$curStats{'afp'}{'neg'}{'ncur'}{$datatype}{'countPap'} = scalar keys %{ $afpNegNC{$datatype} };
$curStats{'afp'}{'neg'}{'ncur'}{$datatype}{'ratio'} = $ratio;
} # foreach my $datatype (@datatypesToShow)
} # sub getCurationStatisticsAfpNeg
"Any" and "Intersection" rows of the Curation Statistics table
To determine the "Any" and "Intersection" results, all flagging methods currently visible in the Curation Statistics table are considered. So, for the main Curation Statistics table (with no options selected), all flagging methods (SVM, AFP, and CFP as of 12-10-2012) are considered. The calculations in this case would be:
Any flagged : %svmData OR %afpFlagged OR %cfpFlagged Any positive : %svmPos OR %afpPos OR %cfpPos Any positive validated : %svmPosVal OR %afpPosVal OR %cfpPosVal Any positive validated false positive : %svmPosFP OR %afpPosFP OR %cfpPosFP Any positive validated true positive : %svmPosTP OR %afpPosTP OR %cfpPosTP Any positive validated true positive curated : %svmPosTpCur OR %afpPosTpCur OR %cfpPosTpCur Any positive validated true positive not curated : %svmPosTpNC OR %afpPosTpNC OR %cfpPosTpNC Any positive not validated : %svmPosNV OR %afpPosNV OR %cfpPosNV Any positive not curated : %svmPosNC OR %afpPosNC OR %cfpPosNC Intersection flagged : %svmData AND %afpFlagged AND %cfpFlagged Intersection positive : %svmPos AND %afpPos AND %cfpPos Intersection positive validated : %svmPosVal AND %afpPosVal AND %cfpPosVal Intersection positive validated false positive : %svmPosFP AND %afpPosFP AND %cfpPosFP Intersection positive validated true positive : %svmPosTP AND %afpPosTP AND %cfpPosTP Intersection positive validated true positive curated : %svmPosTpCur AND %afpPosTpCur AND %cfpPosTpCur Intersection positive validated true positive not curated : %svmPosTpNC AND %afpPosTpNC AND %cfpPosTpNC Intersection positive not validated : %svmPosNV AND %afpPosNV AND %cfpPosNV Intersection positive not curated : %svmPosNC AND %afpPosNC AND %cfpPosNC
Note that if a curator enters the Curation Statistics table after entering deselecting any of the flagging methods in the Curation Statistics Options Page, the "Any" and "Intersection" sections of the table will only reflect the flagging methods chosen by the curator. Thus, if a curator chooses to view only one flagging method, the "Any", "Intersection", and "Flagged Positive" sections of the table will show identical results.
The following are the correspondences between rows in the Curation Statistics table and the hash tables in the form's code:
General paper stats
%curatablePapers curatable papers %objsCurated objects curated %objsCurated/%valCur objects curated per paper %valCur Papers curated %validated Papers validated %valPos Papers validated positive %valCur Papers validated positive curated %valPos NOT %valCur Papers validated positive not curated %valNeg Papers validated negative %conflict Papers validated conflict
Support Vector Machine paper stats
%noSvm SVM no svm processed %svmData SVM has svm %svmPos SVM positive any %svmPosVal SVM positive any validated %svmPosFP SVM positive any validated false positive %svmPosTP SVM positive any validated true positive %svmPosTpCur SVM positive any validated true positive curated %svmPosTpNC SVM positive any validated true positive not curated %svmPosNV SVM positive any not validated %svmPosNC SVM positive any not curated %svmHig SVM positive high %svmHigVal SVM positive high validated %svmHigFP SVM positive high validated false positive %svmHigTP SVM positive high validated true positive %svmHigTpCur SVM positive high validated true positive curated %svmHigTpNC SVM positive high validated true positive not curated %svmHigNV SVM positive high not validated %svmHigNC SVM positive high not curated %svmMed SVM positive medium %svmMedVal SVM positive medium validated %svmMedFP SVM positive medium validated false positive %svmMedTP SVM positive medium validated true positive %svmMedTpCur SVM positive medium validated true positive curated %svmMedTpNC SVM positive medium validated true positive not curated %svmMedNV SVM positive medium not validated %svmMedNC SVM positive medium not curated %svmLow SVM positive low %svmLowVal SVM positive low validated %svmLowFP SVM positive low validated false positive %svmLowTP SVM positive low validated true positive %svmLowTpCur SVM positive low validated true positive curated %svmLowTpNC SVM positive low validated true positive not curated %svmLowNV SVM positive low not validated %svmLowNC SVM positive low not curated %svmNeg SVM negative %svmNegVal SVM negative validated %svmNegTN SVM negative validated true negative %svmNegFN SVM negative validated false negative %svmNegFnCur SVM negative validated false negative curated %svmNegFnNC SVM negative validated false negative not curated %svmNegNV SVM negative not validated %svmNegNC SVM negative not curated
Author First Pass paper stats
%afpEmailed AFP emailed %afpFlagged AFP flagged %afpPos AFP positive %afpPosVal AFP positive validated %afpPosFP AFP positive validated false positive %afpPosTP AFP positive validated true positive %afpPosTpCur AFP positive validated true positive curated %afpPosTpNC AFP positive validated true positive not curated %afpPosNV AFP positive not validated %afpPosNC AFP positive not curated %afpNeg AFP negative %afpNegVal AFP negative validated %afpNegTN AFP negative validated true negative %afpNegFN AFP negative validated false negative %afpNegFnCur AFP negative validated false negative curated %afpNegFnNC AFP negative validated false negative not curated %afpNegNV AFP negative not validated %afpNegNC AFP negative not curated
Curator First Pass paper stats
%cfpFlagged CFP flagged %cfpPos CFP positive %cfpPosVal CFP positive validated %cfpPosFP CFP positive validated false positive %cfpPosTP CFP positive validated true positive %cfpPosTpCur CFP positive validated true positive curated %cfpPosTpNC CFP positive validated true positive not curated %cfpPosNV CFP positive not validated %cfpPosNC CFP positive not curated %cfpNeg CFP negative %cfpNegVal CFP negative validated %cfpNegTN CFP negative validated true negative %cfpNegFN CFP negative validated false negative %cfpNegFnCur CFP negative validated false negative curated %cfpNegFnNC CFP negative validated false negative not curated %cfpNegNV CFP negative not validated %cfpNegNC CFP negative not curated
Abbreviations
AFP - Author First Pass (flagging method)
CFP - Curator First Pass (flagging method)
OA - Ontology Annotator (curation tool)
SVM - Support Vector Machine (flagging method)
Definitions
curated - Any paper that has been curated, as determined by its presence in the OA or in cur_curdata. Note that if a paper is considered 'curated' it is automatically considered 'validated positive'
datatype - A type of data of that WormBase curates
flagged - Processed by a flagging method (flagged positive OR negative)
flagging method - Manual or automated method for identifying research articles that contain a particular datatype
validated - Definitively confirmed by a curator to have (or not have) the relevant datatype