WBConfCall 2014.06.05-Agenda and Minutes
Contents
Agenda and Minutes
New Staff Member Introduction
Sibyl Gao, joining the webdev team at OICR.
- sibyl@wormbase.org
WormBase ParaSite
With ParaSite close to going into production, we should start discussing how to proceed. Topics might include:
- How best to integrate it with the main WormBase site, and with the WormBase release cycle
- The strategy for incorporation of parasitic genomes into WormBase from this point
- Current parasitic species in WormBase - should we "move" them into ParaSite?
- Curation for parasitic species?
This is a big discussion topic, so the intent for this call is to seed some of these discussion, and come up with more concrete plans off-line or in future calls.
WB meeting minutes:
- It is difficult to keep both sites synchronized. Each site can list the dated of last time their data was updated so that users can compare them.
- High value parasite genome (WB core species) should stay in WormBase. It will be confusing to users if we remove them.
- Both sites can link to each other. WB can link to ParaSite for annotation. C. elegans genes on ParaSite gene tree can link back to WB.
- ~300 parasite genomes will be available in the nex couple of years, need suggestions on what to include in gene tree. Paul will visit Hinxton next week and talk with Hinxton group about this.
- Will have another phone conf. about this issue.
nurf-1 Gene structure and naming
In C. elegans, nurf-1 is a complex gene structure that is composed of two main regions. Many isoforms are apparent in this locus. Some of these terminate halfway through the locus, some start in the second half and some span both halves.
In C. briggsae, C. japonica and C. remanei the homologous region to Cel-nurf-1 is composed of two completely separate genes, according to our normal gene curation standards.
In Drosophila and Human, the homologous region is a single complex gene producing many different isoforms, some of which span the two halves, as in C. elegans.
We have a user who is requesting help in naming this locus in C. briggsae for a paper.
There are other examples of complex gene loci that have been annotated in a variety ways. Often our hand has been forced as authors have imposed their view of how these should be named before a discussion of the implications. This has resulted in loci where we have the annotation as:
Existing categories of genes:
Category 1) A single gene locus - single CGC name and single WBGene ID with non-overlapping isoforms.
Category 2) Two completely different genes - two CGC names and two WBGene IDs
Category 3) Two genes with shared Isoform naming - two CGC names and two WBGene IDs - bad and only exists because of users.
Options:
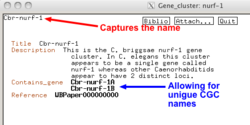
1) Author and Tim: Proposes having a single "locus name" Cbr-nurf-1 having a single CGC name shared between two different genes would work in the database and preserve the current naming but could potentially result in problems for the website. Is this the case? - would be a first so should we discuss nomenclature guidelines?
2) Referee: Proposed nurf-1.1 and nurf-1.2 (Usually reserved for paralogous genes but has been used in the past where a elegans gene has two orthologs in the target species even if they are fragments of the same gene.) - this seems to fit the C. briggsae scenario.
3) lin-15A/B example: Two distinct genes with no shared naming nurf-1A nurf-1B - shows that there is something different in species X compared to species Y - would also fit the C. briggsae scenario.
We could use the Gene_cluster class to store the original name and then have unique cgc names attached to the component genes.
4) Could merge the genes into a single gene with non-overlapping transcript populations....simplifies for website, but not necessarily the best option.
How should we represent and name these sort of loci in the future?
Specific examples of unusual loci for reference:
* Non overlapping transcript populations with a bridging population single gene (like the C. elegans nurf-1) Category 1) A single gene locus - single CGC name and single WBGene ID with non-overlapping isoforms. ------------------------------- unc-105 hpo-12 D2023.1 DH11.5 F11F1.1 - Unusual loci F26H11.2 - nurf-1 * Single transcript population giving rise to 2 gene products ----------------------------------------------------------- ** 1 gene curated Category 1) A single gene locus - single CGC name and single WBGene ID with non-overlapping isoforms. -------------- Prx-10 - Lots of ESTS displaying a single block containing the end and start. ** 2 genes curated Category 2) Two completely different genes - two CGC names and two WBGene IDs --------------- Maf-1::Mgl-2 F32D8.12::F32D8.11 - One appears to be in UTR of other. Nola-3::c25a1.16 - Unusual as nola-3 is very similar to a yeast protein, small and would have been ignored in normal curation Y53C12A.11::Y53C12A.6 - No coding overlap - Lots of ESTS displaying a single block containing the end and start. W01A8.2::W01A8.8 ------------------------------- Non coding transcript populations overlaps 2 sets of coding transcripts also some evidence for a retained intron so could be a single transcript that is processed to give 2 functional forms? Category 3) Two genes with shared Isoform naming - two CGC names and two WBGene IDs * Complex transcript populations with some shared space. lev-10::eat-18 ------------------------------- Y105E8A.7 a/b/e = lev-10 Y105E8A.7 c/d = eat-18 Small amounts of out of frame coding overlap Cha-1::unc-17 ------------------------------- ZC416.8a = unc-17 ZC416.8b = cha-1 No coding overlap just shared transcriptional space in a UTR exon.
WB Meeting Minutes:
- We can call them nurf-1A and nurf-1B, adopt a new class called "Locus" to group them together
- Need to involve Tim Schedle for naming of the genes
Outstanding Help Desk Issues
- Question about other species' data in WormMine - any plans on this front?
Abby: no plan for other species in WormMine. Priority is updating data and include RNAi and Sequence curation.
- Some outstanding GBrowse issues - #2735 and #2745
Abby assigned Todd to respond to users who notified us about the bugs. #2720 and #2744 are for the same bug.