Difference between revisions of "Pictures"
| Line 24: | Line 24: | ||
[[File:PictureA.png]] | [[File:PictureA.png]] | ||
| + | |||
| + | |||
| + | Pictures are saved with their original name in order to minimize editing from the curator. In this case the file is called “journal.pbio.0020352.g006”. | ||
| + | |||
| + | The file is saved in a directory named after the WB paper ID. E.g.: WBPaper00024505, meaning that picture “journal.pbio.0020352.g006” has been downloaded from WBPaper00024505. | ||
| + | |||
| + | |||
| + | [[File:PictureB.png]] | ||
| + | |||
| + | These 2 numbers together WBPaper00024505_journal.pbio.0020352.g006 will be UNIQUE IDENTIFIERS of the object, that we call Picture object 1 (WBPicture000000001). | ||
| + | The path WBPaper00024505_journal.pbio.0020352.g006 will define the SOURCE of the object in the [[picture data model]]. | ||
==Model elements== | ==Model elements== | ||
Revision as of 18:29, 29 September 2010
links to relevant pages
Caltech documentation
PIctures
Contents
Picture Curation
The immediate goal of picture curation is to be able to obtain images of gene expression data from the literature and individual laboratories and display them in the WormBase gene expression page.
- We want display images related to the temporal or spatial (e.g., tissue, subcellular, etc.) localization of any gene in a wild-type background with different data types
- Reporter gene analysis
- Antibody staining
- In situ hybridization
- RT-PCR
- Western or Northern blot data
Pipeline
In the early phases of curation, pictures will be taken from open access journals (e.g. PLoS). During the process of PLoS image curation, other publishers will be contacted for obtaining copyright permissions.
The images should be saved and stored according to the following guidelines. The example shown below refers to a PLoS Biology paper but the rules of handling the pictures are universal and not "paper specific".
- STEP 1
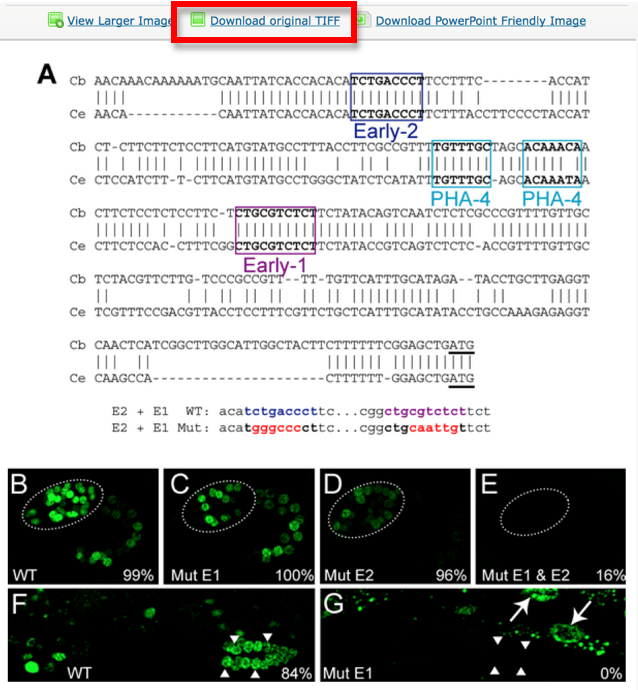
Pictures are downloaded in TIFF format from the original paper.
Pictures are saved with their original name in order to minimize editing from the curator. In this case the file is called “journal.pbio.0020352.g006”.
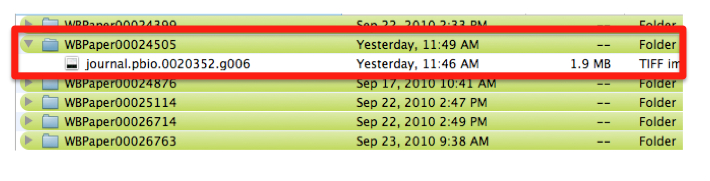
The file is saved in a directory named after the WB paper ID. E.g.: WBPaper00024505, meaning that picture “journal.pbio.0020352.g006” has been downloaded from WBPaper00024505.
These 2 numbers together WBPaper00024505_journal.pbio.0020352.g006 will be UNIQUE IDENTIFIERS of the object, that we call Picture object 1 (WBPicture000000001). The path WBPaper00024505_journal.pbio.0020352.g006 will define the SOURCE of the object in the picture data model.
Model elements
- Name-> MeSH UID
- Public name -> common name in elegans literature
- Synonym -> other names, how do we mine these from other DBs?
- DB_info -> links to entity in other database add following databases to database.ace
Molecule curation
Drug-phenotype curation
Molecules will be linked to genes based on their influence on gene activity altered by variation, overexpression, and RNAi-based knockdown.
Drug-gene interactions
Molecules will also be linked to genes through their influence on gene activity directly through gene regulation interactions.
Molecule databases
Molecule IDs will be provided, when available, for the following databases:
- Database "NLM_MeSH" "UID"
- Database "CTD" "ChemicalID"
- Database "ChemIDplus" using the CasRN
- Database "ChEBI" "CHEBI_ID"
- Database "KEGG COMPOUND" "ACCESSION_NUMBER"
Molecule list
Initially, we will be using MeSH UIDs, assigned by the NLM, as IDs for the molecules in our database. Due to the more comprehensive coverage of the NLM molecules, and the fact that it is more stably funded, this source was thought to be a good starting point for this project. The list we are starting with is a pared down list of molecules from the NLM, that was created by the Comparative Toxicogenomic Database (CTD), which contains over 130,000 terms. For each term, this list contains a term name, CTD ID, MeSH UID, and where available CAS Registry Numbers. Using the CasRNs, we extracted the ChEBI ID from the Chemical Entities of Biological Interest database entity list, where it existed, along with any KEGG Compound accession number.
A sample molecule.ace record:
Molecule : "C009687" Public_name "wortmannin" Database "NLM_MeSH" "UID" "C009687" Database "CTD" "ChemicalID" "C009687" Database "ChemIDplus" "19545-26-7" Database "ChEBI" "CHEBI_ID" "52289" Database "KEGG COMPOUND" "ACCESSION_NUMBER" "C15181"