Phenotype micropublications
For Phenotype-micropublications submissions we will modify the existing Allele-phenotype submission form. The underlying CGI will remain the same but the Allele-phenotype and the Phenotype-micropublications form will have 2 separate displays.
Contents
Changes to the form
Changes to the current Allele-phenotype form located here: http://www.wormbase.org/submissions/phenotype.cgi: Expression micropublication form here: http://mangolassi.caltech.edu/~azurebrd/cgi-bin/forms/expr_micropub.cgi
Change the descriptive text at the beginning with:
'If you have unpublished phenotype data you can micropublish it on WormBase by filling the form below. The form is mostly self explanatory and hints can be found by clicking green question marks. In the box on the right side of the page you can find additional tips/information that will appear when you start typing in specific fields. Should you have additional questions you can find guidelines here.' Remove:
- PubMedID -submissions are novel data, do not have a paper attached. Submissions will get their own WBPaperID
- Click here to review your publications and see which are in need of phenotype curation -this field not necessary in the micropublication form
Add -below 'Your e-mail address' field:
- co-authors -> autocomplete on people
- Laboratory -> autocomplete on labs -> mandatory
- Funding -> free text -> mandatory
- Species -> autocomplete on species?
- Choose an image -> Browse button, only jpg accepted -> mandatory
- Description -> big text (small box that becomes big upon clicking)
Change:
- Genetic perturbation(s) in -> Perturbation(s)
- this text: 'PLEASE NOTE: All genetic perturbations above will be annotated to all phenotypes entered below. For separate perturbation-phenotype annotations, please perform separate submissions, or use the WormBase Phenotype Worksheet (linked above).'
in this: 'PLEASE NOTE: All perturbations above will be annotated to all phenotypes entered below. For separate perturbation-phenotype annotations, please perform separate submissions, or use the WormBase Phenotype Worksheet (linked above).'
Add -below 'Transgene' field: Molecule -> autocomplete on molecules
Add -above 'Optional'
- Disclaimer -> mandatory checkbox like in the Expression micropublication form: I declare to the best of my knowledge that the experiment is reproducible.
Model Changes
RNAi
in ?RNAi class add: Supporting_data Picture ?Picture XREF RNAi
in ?Picture class add RNAi ?RNAi XREF Picture
#Phenotype_info
In #Phenotype_info add Picture ?Picture (can we put Xrefs to the hash? XREF #Phenotype_info)
In ?Picture add ---
Changes to the OAs
RNAi OA
Add a Picture field in tab 4 between 'History name' and 'Movie'
Phenotype OA
Add a Picture field
Title?
Shall we add a 'Title' field or shall we autogenerate titles with the info that are entered in different fields?
Modelling Nemametrix data
notes
The strain is a model for human disease Can the annotation be to strain-phenotype?
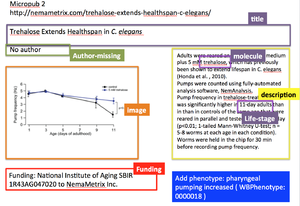
No funding listed for this experiment on the nemametrix site Add: Funding: National Institute of Aging SBIR 1R43AG047020 to NemaMetrix Inc.
What is listed as figure legend 1 should go into reagents- also other tech notes do not have the caption but only the descriptive text