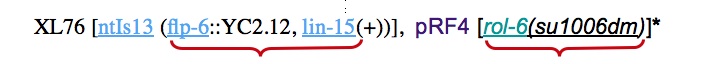Markup excerpts
From WormBaseWiki
Jump to navigationJump to searchback to GSA markup SOP
back to Entity classes and problems
Marked up html index
Classes and problems
Anatomy_name
This is an example of text markup testing the feasibility of linking based on anatomy_name. As you will see, there are a number of problems with the links that are created.
| Anatomy_name | Problem | Possible solution |
|---|---|---|
| “set”, “cell” | Real term but also common word | Make context specific |
| “CAN” | Real term but also common word | Use case sensitivity |
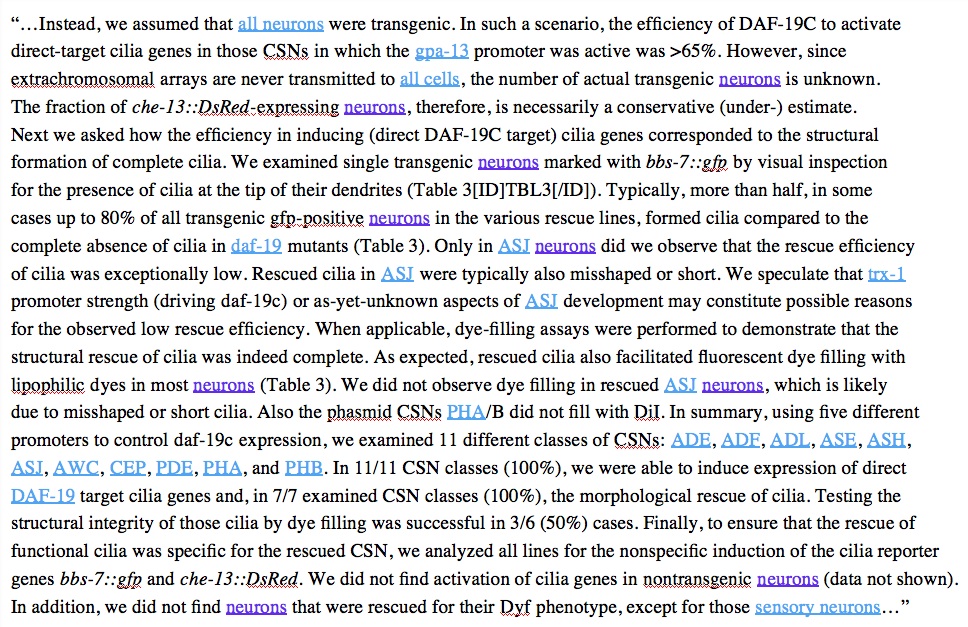
| “AC”, “FLP” | Real term but also different meanings “… (<AC>pAC> < 0.001) and 0.95 (<AC>pAC> < 0.001) in wild-type” AC is not anchor cell “e.g., the FLP recombinase system (Davis et al. 2008).” FLP is not neuron cell | Manual correction…context specificity? |
| “organism”,“organ”, “axis”, “neuronal”, “head”, tail” | Too general for the researcher? | Remove from list? |
| “PhaA/B”, "CAN", "VPC" | Synonyms not recognized: “CSN” for ciliated sensory neuron, “VPC”s for vulval precursor cells, “PhaA/B” | Add to entity list |
| “neuron” okay, “neurons” not okay | Plurality problems | Make plural indifferent |
| “Lateral sense organ”, “P12 ectoblast” not linked | Term not on list: "P12" ok but the expression "P12 ectoblast" isn’t, Synonym problem?? | Add term? Add synonym? Just omit for linking? |
| Single letters “D”, “K”, etc | Too many false hits: Protein domains “C terminus”, “D domain", Figure panel references “in panel A of Figure 1” | Remove single letters from list, or make context specific. |
Genomic Expressions
Genomic expressions are not correctly linked
It is not correct to link transgene expressions to the individual entities within the construct
*Note: This genotype is hidden from web page in the tag General_remark, same as for other plasmids ex. HSP-FLP1, pRF4, cpjM1HG.11, IM#171, p76-16B, pPD118.33, pTG96, etc.
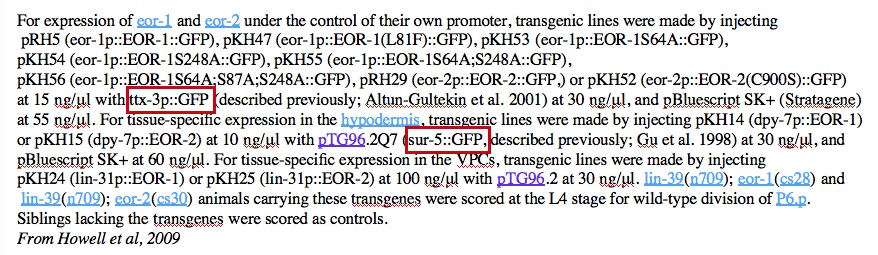
- Common name is genotype, not actual name:
- ttx-3p::GFP, in this case, the transgene is mgIs18, but not stated as such in the paper. The expression “[ttx-3::GFP]” in the Summary of mgIs18 in acedb
- sur-5::GFP
- Solution:
- all gene links within genomic expressions will be unmade