Difference between revisions of "Gene Interaction/archive"
m |
m (→Old Models) |
||
| (16 intermediate revisions by the same user not shown) | |||
| Line 36: | Line 36: | ||
===Old Models=== | ===Old Models=== | ||
| + | |||
| + | Updated for WS249 - '''CG 4-2-2015''' | ||
| + | |||
| + | <pre> | ||
| + | ?Interaction Interaction_type Physical | ||
| + | Predicted | ||
| + | Regulatory Change_of_localization | ||
| + | Change_of_expression_level | ||
| + | Genetic Genetic_interaction | ||
| + | Neutral_genetic | ||
| + | Synthetic | ||
| + | Enhancement | ||
| + | Unilateral_enhancement | ||
| + | Mutual_enhancement | ||
| + | Positive_genetic | ||
| + | Suppression | ||
| + | Complete_suppression | ||
| + | Partial_suppression | ||
| + | Unilateral_suppression | ||
| + | Complete_unilateral_suppression | ||
| + | Partial_unilateral_suppression | ||
| + | Mutual_suppression | ||
| + | Complete_mutual_suppression | ||
| + | Partial_mutual_suppression | ||
| + | Suppression_enhancement | ||
| + | Asynthetic | ||
| + | Negative_genetic | ||
| + | Oversuppression | ||
| + | Unilateral_oversuppression | ||
| + | Mutual_oversuppression | ||
| + | Oversuppression_enhancement | ||
| + | Phenotype_bias | ||
| + | No_interaction // Negative data; no interaction was observed after testing | ||
| + | Epistasis | ||
| + | Positive_epistasis | ||
| + | Maximal_epistasis | ||
| + | Minimal_epistasis | ||
| + | Neutral_epistasis | ||
| + | Qualitative_epistasis | ||
| + | Opposing_epistasis | ||
| + | Quantitative_epistasis | ||
| + | Interactor PCR_interactor ?PCR_product XREF Interaction #Interactor_info // PCR_product of the interacting gene or protein, e.g. Yeast Two Hybrid experiments | ||
| + | Sequence_interactor ?Sequence XREF Interaction #Interactor_info // Sequence of the interacting gene or protein | ||
| + | Feature_interactor ?Feature XREF Associated_with_Interaction #Interactor_info | ||
| + | Interactor_overlapping_CDS ?CDS XREF Interaction #Interactor_info // CDS of the interacting gene or protein (or related sequence) | ||
| + | Interactor_overlapping_gene ?Gene XREF Interaction #Interactor_info // Gene (or portion of gene) involved in the interaction | ||
| + | Interactor_overlapping_protein ?Protein XREF Interaction #Interactor_info // Protein (or portion of protein) involved in the interaction | ||
| + | Molecule_interactor ?Molecule XREF Interaction #Interactor_info // Molecule that interacts with a gene or protein (ported from Gene_regulation Class) | ||
| + | Other_interactor ?Text #Interactor_info // Free text describing an interactor entity or condition that does not fall into a standard WormBase category | ||
| + | Rearrangement ?Rearrangement XREF Interactor #Interactor_info | ||
| + | Variation_interactor ?Variation XREF Interactor #Interactor_info // Added WS248; allele involved in genetic interaction | ||
| + | Interaction_summary ?Text #Evidence | ||
| + | Detection_method Affinity_capture_luminescence // A physical interaction detection technique | ||
| + | Affinity_capture_MS // A physical interaction detection technique | ||
| + | Affinity_capture_RNA // A physical interaction detection technique | ||
| + | Affinity_capture_Western // A physical interaction detection technique | ||
| + | Chromatin_immunoprecipitation // A physical interaction detection technique | ||
| + | Cofractionation // A physical interaction detection technique | ||
| + | Colocalization // A physical interaction detection technique | ||
| + | Copurification // A physical interaction detection technique | ||
| + | DNase_I_footprinting // A physical interaction detection technique | ||
| + | Fluorescence_resonance_energy_transfer // A physical interaction detection technique | ||
| + | Protein_fragment_complementation_assay // A physical interaction detection technique | ||
| + | Yeast_two_hybrid // A physical interaction detection technique (Protein-protein) | ||
| + | Biochemical_activity // A physical interaction detection technique | ||
| + | Cocrystal_structure // A physical interaction detection technique | ||
| + | Far_western // A physical interaction detection technique | ||
| + | Protein_peptide // A physical interaction detection technique | ||
| + | Protein_RNA // A physical interaction detection technique | ||
| + | Reconstituted_complex // A physical interaction detection technique | ||
| + | Electrophoretic_mobility_shift_assay // A physical interaction detection technique | ||
| + | Yeast_one_hybrid // A physical interaction detection technique (Protein-DNA) | ||
| + | Directed_yeast_one_hybrid // A physical interaction detection technique (Protein-DNA) | ||
| + | Antibody // A regulatory interaction detection technique; Antibody name and details captured in Interactor_info hash | ||
| + | Reporter_gene ?Text // A regulatory interaction detection technique | ||
| + | Transgene // A regulatory interaction detection technique; Trasnsgene name and details captured in Interactor_info hash | ||
| + | In_situ Text // A regulatory interaction detection technique | ||
| + | Northern Text // A regulatory interaction detection technique | ||
| + | Western Text // A regulatory interaction detection technique | ||
| + | RT_PCR Text // A regulatory interaction detection technique | ||
| + | Other_method ?Text // A regulatory interaction detection technique | ||
| + | Library_info Library_screened Text INT // In the context of Y2H or YH screens, for example, the library may have been cDNA library or a pool of clones | ||
| + | Origin From_laboratory UNIQUE ?Laboratory // A library generated at an academic laboratory | ||
| + | From_company UNIQUE ?Text // A library generated at a company | ||
| + | Regulation_level Transcriptional // Regulation occurs at the transcriptional level | ||
| + | Post_transcriptional // Regulation occurs at the post-transcriptional level | ||
| + | Post_translational // Regulation occurs at the post-translational level | ||
| + | Regulation_result Positive_regulate #GR_condition | ||
| + | Negative_regulate #GR_condition | ||
| + | Does_not_regulate #GR_condition // added to capture negative data [040220 krb] | ||
| + | Confidence Description Text // Free text description of the confidence, e.g. "Core" vs "Noncore" (Vidal Interactome terms) | ||
| + | P_value UNIQUE Float // P-value confidence of interaction, if given | ||
| + | Log_likelihood_score UNIQUE Float // Only used for Predicted interactions | ||
| + | Throughput UNIQUE High_throughput //See BioGRID curation criteria for discussion | ||
| + | Low_throughput | ||
| + | Interaction_RNAi ?RNAi XREF Interaction // RNAi experiment associated with the interaction | ||
| + | Interaction_phenotype ?Phenotype XREF Interaction // Phenotype associated with a genetic interaction | ||
| + | Unaffiliated_variation ?Variation | ||
| + | Unaffiliated_transgene ?Transgene | ||
| + | Unaffiliated_antibody ?Antibody | ||
| + | Unaffiliated_expr_pattern ?Expr_pattern | ||
| + | Unaffiliated_construct ?Construct | ||
| + | WBProcess ?WBProcess XREF Interaction // WormBase biological process associated with the interaction | ||
| + | DB_info Database ?Database ?Database_field ?Text // Any database reference to the interaction outside of WormBase, e.g. BioGRID, Interactome | ||
| + | Paper ?Paper XREF Interaction | ||
| + | Antibody_remark ?Text | ||
| + | Historical_gene ?Gene Text | ||
| + | Remark ?Text #Evidence | ||
| + | |||
| + | </pre> | ||
| Line 87: | Line 197: | ||
Mutual_suprression//non_directional | Mutual_suprression//non_directional | ||
/////////////////////////////////////////////////////////////////////////////////// | /////////////////////////////////////////////////////////////////////////////////// | ||
| + | |||
| + | ==== Tags removed for WS248 ==== | ||
| + | |||
| + | From ?Interaction model | ||
| + | |||
| + | <pre> | ||
| + | Deviation_from_expectation Text // A text description of the way in which the phenotype deviated from expectation in genetic interactions | ||
| + | Neutrality_function UNIQUE Multiplicative // The multiplicative neutrality function defines expectation as the product of two quantified phenotypes (relative to wild type) | ||
| + | Additive // The additive neutrality function defines expectation as the sum of two quantified phenotypes (relative to wild type) | ||
| + | Minimal // The minimal neutrality function defines expectation as the most severe of two quantified phenotypes (relative to wild type) | ||
| + | </pre> | ||
| + | |||
| + | From #Interactor_info model | ||
| + | |||
| + | <pre> | ||
| + | Intragenic_effector_variation ?Variation XREF Interactor | ||
| + | Intragenic_affected_variation ?Variation XREF Interactor | ||
| + | Variation ?Variation XREF Interactor // (allele, polymorphism, etc.) involved in the interaction //removed XREF from proposal | ||
| + | </pre> | ||
===New Model Proposals=== | ===New Model Proposals=== | ||
| Line 434: | Line 563: | ||
'''Old Table Names''' | '''Old Table Names''' | ||
| − | |||
'Non_directional' = int_nondirectional | 'Non_directional' = int_nondirectional | ||
| Line 446: | Line 574: | ||
'Affected Transgene Gene' = int_transgenetwogene | 'Affected Transgene Gene' = int_transgenetwogene | ||
| − | < | + | 'Deviation from expectation' = int_deviation |
| + | |||
| + | 'Neutrality function' = int_neutralityfxn | ||
| + | |||
| + | 'Intragenic Effector Variation(s)' = int_intravariationone | ||
| + | |||
| + | 'Intragenic Affected Variation(s)' = int_intravariationtwo | ||
| + | |||
| + | 'Effector Other Type' = int_otheronetype | ||
| + | |||
| + | 'Affected Other Type' = int_othertwotype | ||
| + | |||
| + | === Removed OA fields === | ||
| + | * As of 2-19-2015 | ||
| + | |||
| + | ==== TAB3 ==== | ||
| + | *Intragenic Effector Variation(s) - ?Variation (MultiOntology) '''int_intravariationone''' | ||
| + | **'''Genes for this field need to be mapped to a gene at the dump stage; Genes that map to the variation will be dumped as the "Interactor_overlapping_gene" as follows: | ||
| + | **'''Dumps as: Interactor_overlapping_gene <Mapped Gene> Intragenic_effector_variation <Variation>''' | ||
| + | **'''For Variations that don't map to a gene in the interaction, Dump as: Unaffiliated_variation <Variation>''' | ||
| + | |||
| + | *Intragenic Affected Variation(s) - ?Variation (MultiOntology) '''int_intravariationtwo''' | ||
| + | **'''Genes for this field need to be mapped to a gene at the dump stage; Genes that map to the variation will be dumped as the "Interactor_overlapping_gene" as follows: | ||
| + | **'''Dumps as: Interactor_overlapping_gene <Mapped Gene> Intragenic_affected_variation <Variation>''' | ||
| + | **'''For Variations that don't map to a gene in the interaction, Dump as: Unaffiliated_variation <Variation>''' | ||
| + | |||
| + | |||
| + | ==== TAB4 ==== | ||
| + | *Deviation from expectation - Big text '''int_deviation''' '''Dumps as: Deviation_from_expectation <Big_Text>''' | ||
| + | |||
| + | *Neutrality function - (Dropdown) '''int_neutralityfxn''' options are: | ||
| + | **Multiplicative '''Dumps as: Multiplicative''' | ||
| + | **Additive '''Dumps as: Additive''' | ||
| + | **Minimal '''Dumps as: Minimal''' | ||
| + | |||
| + | *Effector Other Type - (Dropdown) int_otheronetype options are: Chemical or Transgene, int_otheronetype '''Dumps as (see next line)''' | ||
| + | |||
| + | *Affected Other Type - (Dropdown) int_othertwotype options are: Chemical or Transgene, int_othertwotype '''Dumps as (see next line)''' | ||
| + | |||
| + | How to use: | ||
| + | |||
| + | '''Deviation from expectation''' is a free-text field where, at the curator's discretion, a curator can describe why a genetic interaction is an interaction, i.e. why it is an unexpected result warranting an interaction. This could be as simple as "The life spans were synergistic" or "Neither mutation alone exhibited a phenotype". | ||
| + | '''Neutrality function''' is closely related to the "Deviation from expectation" field, where the curator can decide, if applicable, which "Neutrality function" applies to this genetic interaction. The choices in this drop down field are "Multiplicative", "Additive", and "Minimal". "Multiplicative" means that the authors expected to see a quantifiable phenotype in the double mutant that was the mathematical product of the quantified phenotypes of the individual mutations. For example, one mutant extends life span by 20%, and another extends it by 30%. With a multiplicative neutrality function, we would expect the double mutant to have (1.2 * 1.3 = 1.56) about 56% extended life span. Alternatively, in the "Additive" neutrality function, the authors might expect the sum of the effects (A + B - 1) or (1.2 + 1.3 - 1 = 1.5) or a 50% extended life span. The "Minimal" neutrality function assumes that the double mutant will be as severe as the most severe single mutant, in this case (1.3) or 30% extended. Therefore, any life span extension beyond 30%, for example, in the double mutant would be considered "surprising" and therefore, an interaction. | ||
| + | '''Effector/Affected Other Type''' and '''Effector/Affected Other''' fields: these fields allow for the curation of Chemicals, Transgenes, or other entities that don't exists as proper WormBase/ACEDB objects, for example transgenes that only express human proteins. The "Other Type" fields allow for the selection of "Chemical" or "Transgene", the identity of which would go into the "Other" fields. This, ideally, will get phased out as chemicals are generated in the Molecule OA and transgenes in the Transgene OA, thereby allowing them to be entered into ontology-based "Transgene" or "Molecule" fields. | ||
==='''.ace template: (OUTDATED)'''=== | ==='''.ace template: (OUTDATED)'''=== | ||
| Line 520: | Line 691: | ||
==='''interaction objects source file'''=== | ==='''interaction objects source file'''=== | ||
| − | + | ||
*there are 9242 interaction objects dumped from WS220 on Monday, 10/01/2010 | *there are 9242 interaction objects dumped from WS220 on Monday, 10/01/2010 | ||
| Line 537: | Line 708: | ||
Also there are interaction data in postgres without an interaction ID//will be assigned id from WBInteraction0500001. | Also there are interaction data in postgres without an interaction ID//will be assigned id from WBInteraction0500001. | ||
| + | |||
| + | == Some Notes for gene_gene_interaction == | ||
| + | |||
| + | ==== Outdated notes on large scale interaction files ==== | ||
| + | *<strike> ready_to_upload large scale data file: Large_scale_interactions_WS231.ace is located on tazendra: /home/acedb/xiaodong/oa_interactions_dumper should be uploaded for every upload | ||
| + | * large scale .ace file needs to be checkd again dead genes for each upload using scripts in the same directory: find_invalid_genes_largescale.pl* - 08/31/2012 for upload WS234 | ||
| + | **before run, change source file (in file) to latest large scale .ace version (in scripts) | ||
| + | **dead/merged genes will appear on screen. replace merged genes, leave dead/dead genes for now | ||
| + | **change large scale.ace file name to latest upload version | ||
| + | **then upload </strike> | ||
| + | |||
| + | |||
| + | === Citace upload notes === | ||
| + | *2011.1.27 | ||
| + | **caught-up at acedb reading: WBInteraction0500069 (Karen), new line in remark field. fixed in .ace file. | ||
| + | **some confusion on ids. found out ticket issuer was using sandbox data. fixed. | ||
| + | *2011.5.4 | ||
| + | **Karen needed to update variation_wbgene file on tazendra: /home/acedb/jolene/WS_AQL_queries/Variation_gene.txt | ||
| + | **Juancarlos changed the dumper to ignore line breaks and double spaces in remark field for dumping ace file | ||
Latest revision as of 15:10, 21 July 2017
Contents
- 1 Interaction Curation
- 1.1 Pipeline
- 1.2 Some Notes for gene_gene_interaction
Interaction Curation
Pipeline
semi-automatic curation with textpresso extracted sentences
- There are 2138 sentences (actually 2133 sentences) in the sourcefile /home/postgres/work/pgpopulation/genegeneinteraction/20091002-xiaodong/ggi_20091002
- paper starts at WBPaper00028425, ends at WBPaper00035225
- .ace dumper at /home/acedb/xiaodong/gene_gene_interaction/dump_ggi_ace.pl
- go to the directory and do: ./dump_ggi_ace.pl > some_file.ace
- Populate textpresso data in tazendra OA: done on 20110110 -X**
- cd to directory on tazendra: /home/acedb/xiaodong/textpresso_ggi
- mkdir directionay_name (eg 20110106)
- cd directory_name (eg 20110106)
- get Arun's result file (35225-35725.txt in the directory)
- run script: ./populate_textpresso_ggi_to_OA.pl 20110106/35225-35725.txt WBPerson4793 > 20110106/35225-35725.pg (with first argument file_name as input file, and second argument WBPersonID, then output file)
- after running, '20110106/35225-35725.pg' should be in '20111106' directory.
upload Gary and Chris RNAi-based interaction objects into OA
- Reading file created by Igor's script into aceDB
- you can use the empty database by ssh -X citpub@spica.caltech.edu
- then cd CitaceMirror
- then type 'ts' to launch an empty acedb
- Dumping no-worry .ace file
- Then parse into OA
- scp ace file to same directory as below
- ssh acedb@tazendra.caltech.edu (directory xiaodong/interaction_ace_parsing)
- To run : ./parse_ace_interaction_oa.pl gary_RNAi.ace WBPerson1823
- where gary_RNAi.ace is the <inputfile>,the first argument, and WBPerson# is the second argument.
Old Models
Updated for WS249 - CG 4-2-2015
?Interaction Interaction_type Physical
Predicted
Regulatory Change_of_localization
Change_of_expression_level
Genetic Genetic_interaction
Neutral_genetic
Synthetic
Enhancement
Unilateral_enhancement
Mutual_enhancement
Positive_genetic
Suppression
Complete_suppression
Partial_suppression
Unilateral_suppression
Complete_unilateral_suppression
Partial_unilateral_suppression
Mutual_suppression
Complete_mutual_suppression
Partial_mutual_suppression
Suppression_enhancement
Asynthetic
Negative_genetic
Oversuppression
Unilateral_oversuppression
Mutual_oversuppression
Oversuppression_enhancement
Phenotype_bias
No_interaction // Negative data; no interaction was observed after testing
Epistasis
Positive_epistasis
Maximal_epistasis
Minimal_epistasis
Neutral_epistasis
Qualitative_epistasis
Opposing_epistasis
Quantitative_epistasis
Interactor PCR_interactor ?PCR_product XREF Interaction #Interactor_info // PCR_product of the interacting gene or protein, e.g. Yeast Two Hybrid experiments
Sequence_interactor ?Sequence XREF Interaction #Interactor_info // Sequence of the interacting gene or protein
Feature_interactor ?Feature XREF Associated_with_Interaction #Interactor_info
Interactor_overlapping_CDS ?CDS XREF Interaction #Interactor_info // CDS of the interacting gene or protein (or related sequence)
Interactor_overlapping_gene ?Gene XREF Interaction #Interactor_info // Gene (or portion of gene) involved in the interaction
Interactor_overlapping_protein ?Protein XREF Interaction #Interactor_info // Protein (or portion of protein) involved in the interaction
Molecule_interactor ?Molecule XREF Interaction #Interactor_info // Molecule that interacts with a gene or protein (ported from Gene_regulation Class)
Other_interactor ?Text #Interactor_info // Free text describing an interactor entity or condition that does not fall into a standard WormBase category
Rearrangement ?Rearrangement XREF Interactor #Interactor_info
Variation_interactor ?Variation XREF Interactor #Interactor_info // Added WS248; allele involved in genetic interaction
Interaction_summary ?Text #Evidence
Detection_method Affinity_capture_luminescence // A physical interaction detection technique
Affinity_capture_MS // A physical interaction detection technique
Affinity_capture_RNA // A physical interaction detection technique
Affinity_capture_Western // A physical interaction detection technique
Chromatin_immunoprecipitation // A physical interaction detection technique
Cofractionation // A physical interaction detection technique
Colocalization // A physical interaction detection technique
Copurification // A physical interaction detection technique
DNase_I_footprinting // A physical interaction detection technique
Fluorescence_resonance_energy_transfer // A physical interaction detection technique
Protein_fragment_complementation_assay // A physical interaction detection technique
Yeast_two_hybrid // A physical interaction detection technique (Protein-protein)
Biochemical_activity // A physical interaction detection technique
Cocrystal_structure // A physical interaction detection technique
Far_western // A physical interaction detection technique
Protein_peptide // A physical interaction detection technique
Protein_RNA // A physical interaction detection technique
Reconstituted_complex // A physical interaction detection technique
Electrophoretic_mobility_shift_assay // A physical interaction detection technique
Yeast_one_hybrid // A physical interaction detection technique (Protein-DNA)
Directed_yeast_one_hybrid // A physical interaction detection technique (Protein-DNA)
Antibody // A regulatory interaction detection technique; Antibody name and details captured in Interactor_info hash
Reporter_gene ?Text // A regulatory interaction detection technique
Transgene // A regulatory interaction detection technique; Trasnsgene name and details captured in Interactor_info hash
In_situ Text // A regulatory interaction detection technique
Northern Text // A regulatory interaction detection technique
Western Text // A regulatory interaction detection technique
RT_PCR Text // A regulatory interaction detection technique
Other_method ?Text // A regulatory interaction detection technique
Library_info Library_screened Text INT // In the context of Y2H or YH screens, for example, the library may have been cDNA library or a pool of clones
Origin From_laboratory UNIQUE ?Laboratory // A library generated at an academic laboratory
From_company UNIQUE ?Text // A library generated at a company
Regulation_level Transcriptional // Regulation occurs at the transcriptional level
Post_transcriptional // Regulation occurs at the post-transcriptional level
Post_translational // Regulation occurs at the post-translational level
Regulation_result Positive_regulate #GR_condition
Negative_regulate #GR_condition
Does_not_regulate #GR_condition // added to capture negative data [040220 krb]
Confidence Description Text // Free text description of the confidence, e.g. "Core" vs "Noncore" (Vidal Interactome terms)
P_value UNIQUE Float // P-value confidence of interaction, if given
Log_likelihood_score UNIQUE Float // Only used for Predicted interactions
Throughput UNIQUE High_throughput //See BioGRID curation criteria for discussion
Low_throughput
Interaction_RNAi ?RNAi XREF Interaction // RNAi experiment associated with the interaction
Interaction_phenotype ?Phenotype XREF Interaction // Phenotype associated with a genetic interaction
Unaffiliated_variation ?Variation
Unaffiliated_transgene ?Transgene
Unaffiliated_antibody ?Antibody
Unaffiliated_expr_pattern ?Expr_pattern
Unaffiliated_construct ?Construct
WBProcess ?WBProcess XREF Interaction // WormBase biological process associated with the interaction
DB_info Database ?Database ?Database_field ?Text // Any database reference to the interaction outside of WormBase, e.g. BioGRID, Interactome
Paper ?Paper XREF Interaction
Antibody_remark ?Text
Historical_gene ?Gene Text
Remark ?Text #Evidence
?Interaction Evidence #Evidence
Interactor ?Gene XREF Interaction #Interactor_info
Interaction_type Genetic #Interaction_info
Regulatory #Interaction_info
No_interaction #Interaction_info
Predicted_interaction #Interaction_info
Physical_interaction #Interaction_info
Suppression #Interaction_info
Enhancement #Interaction_info
Synthetic #Interaction_info
Epistasis #Interaction_info
Mutual_enhancement #Interaction_info
Mutual_suppression #Interaction_info
Confidence Confidence_level UNIQUE Float
P_value UNIQUE Float
Log_likelihood_score UNIQUE Float
Paper ?Paper XREF Interaction
DB_info Database ?Database ?Database_field ?Accession_number
Remark ?Text #Evidence
#Interactor_info
Variation ?Variation XREF Interactor
Transgene ?Transgene XREF Interactor
Remark ?Text #Evidence //info about the reagents that the model can't capture goes here (e.g. co_suppression, RNA_reagent, etc.)
#Interaction_info
Interaction_RNAi ?RNAi XREF Interaction
Effector ?Gene //master, upstream
Effected ?Gene //subject, downstream
Non_directional ?Gene //e.g. synthetic interactions - Igor
Interaction_phenotype ?Phenotype XREF Interaction
Confidence Confidence_level UNIQUE Float
P_value UNIQUE Float
#Interaction_type
Genetic //directional and non_directional
Physical_interaction
Regulation
No_interaction
Synthetic//non_directional
Epistasis
Enhancement
Suppression
Predicted //addition for WeiWei, non_direactional
Mutual_enhancement//non_directional
Mutual_suprression//non_directional
///////////////////////////////////////////////////////////////////////////////////
Tags removed for WS248
From ?Interaction model
Deviation_from_expectation Text // A text description of the way in which the phenotype deviated from expectation in genetic interactions
Neutrality_function UNIQUE Multiplicative // The multiplicative neutrality function defines expectation as the product of two quantified phenotypes (relative to wild type)
Additive // The additive neutrality function defines expectation as the sum of two quantified phenotypes (relative to wild type)
Minimal // The minimal neutrality function defines expectation as the most severe of two quantified phenotypes (relative to wild type)
From #Interactor_info model
Intragenic_effector_variation ?Variation XREF Interactor Intragenic_affected_variation ?Variation XREF Interactor Variation ?Variation XREF Interactor // (allele, polymorphism, etc.) involved in the interaction //removed XREF from proposal
New Model Proposals
Physical Interaction Model v1.3
?Physical_interaction Evidence #Evidence
Interactor Non_directional_interactor PCR_non_directional_interactor UNIQUE ?PCR_product XREF to? ?Species
Sequence_non_directional_interactor UNIQUE ?Sequence XREF to? ?Species
Non_directional_interactor_overlapping_CDS ?CDS XREF to? ?Species #Evidence
Non_directional_interactor_overlapping_gene ?Gene XREF to ? ?Species #Evidence
Non_directional_interactor_DB_info ?Database ?Database_field UNIQUE ?Accession_number //BioGRID, BioGRIDID, Numerical Value
Bait PCR_bait UNIQUE ?PCR_product XREF to? ?Species
Sequence_bait UNIQUE ?Sequence XREF to? ?Species
Bait_overlapping_CDS ?CDS XREF to? ?Species #Evidence
Bait_overlapping_gene ?Gene XREF to? ?Species #Evidence
Bait_DB_info ?Database ?Database_field UNIQUE ?Accession_number //BioGRID, BioGRIDID, Numerical Value
Target PCR_target UNIQUE ?PCR_product XREF to? ?Species
Sequence_target UNIQUE ?Sequence XREF to? ?Species
Target_overlapping_CDS ?CDS XREF to? ?Species #Evidence
Target_overlapping_gene ?Gene XREF to? ?Species #Evidence
Target_DB_info ?Database ?Database_field UNIQUE ?Accession_number //BioGRID, BioGRIDID, Numerical Value
Experimental_System UNIQUE Affinity_capture-luminescence //Experimental_system includes WormBase tags values as well as BioGRID values
Affinity_capture-MS
Affinity_capture-RNA
Affinity_capture-Western
Co-fractionation
Co-localization
Co-purification
FRET
PCA
Two-hybrid
Biochemical_activity
Co-crystal_structure
Far_western
Protein_peptide
Protein_RNA
Reconstituted_complex
Y1H //BioGRID is not curating protein-DNA interactions. WB has both Y1H data and GO MF data.
Directed_Y1H Text
Protein_DNA
Throughput UNIQUE High_throughput //See BioGRID curation criteria for discussion: http://www.yeastgenome.org/help/BiogridCuration.html
Low_throughput
Library_info Library_screened UNIQUE ?Library //This could also just be ?Text. Doesn't look like ?Library class is used.
Origin From_laboratory UNIQUE ?Laboratory //XREF by making a Reagents tag in the ?Laboratory model?
From_company UNIQUE ?Text //We don't currently have a ?Company class. Should we?
Confidence ?Text //Not currently captured by BioGRID, but this tag can accommodate the legacy YH data.
Paper ?Paper XREF to ?
Remark ?Text #Evidence //How would remarks coming from BioGRID be attributed? Person_evidence? Curator_confirmed? Accession_evidence? Person or Curator would require a change to the dumping file from BioGRID.
Physical Interaction Model v1.2
The revised v1 includes: 1) a ?Species tag, 2) a slot to capture Non-directional interactors (for curating things like protein complexes purified over sedimentation gradients, i.e. where there is no clear Bait or Target directionality), and 3) change ?Confidence from a specific list of phrases or statistical methods to a ?Text tag since this information is expressed in many different ways in the literature so including specific text here doesn't seem practical. If we change this to ?Text, then I'd also remove the specific Interactome_core tag.
Also, current XREF tags in the ?YH model are YH_bait and YH_target. What would be a more appropriate name? Model below has Interaction_target, etc. but I think that's not clear enough. What about Physical_interaction_target?
Also, CDS and Gene, when overlapping, have #Evidence, but the PCR and Sequence do not. Why is this? Does it have to do with needing to indicate how a CDS or Gene was selected without a corresponding sequence?
?Physical_interaction Evidence #Evidence
Species UNIQUE ?Species
Interactor Non_directional_interactor PCR_non_directional_interactor UNIQUE ?PCR_product XREF ?
Sequence_non_directional_interactor UNIQUE ?Sequence XREF ?
Non_directional_interactor_overlapping_CDS ?CDS XREF ? #Evidence
Non_directional_interactor_overlapping_gene ?Gene XREF ? #Evidence
Bait PCR_bait UNIQUE ?PCR_product XREF ?
Sequence_bait UNIQUE ?Sequence XREF ?
Bait_overlapping_CDS ?CDS XREF ? #Evidence
Bait_overlapping_gene ?Gene XREF ? #Evidence
Target PCR_target UNIQUE ?PCR_product XREF ?
Sequence_target UNIQUE ?Sequence XREF ?
Target_overlapping_CDS ?CDS XREF ? #Evidence
Target_overlapping_gene ?Gene XREF ? #Evidence
Experiment_type Affinity_capture-luminescence
Affinity_capture-MS
Affinity_capture-RNA
Affinity_capture-Western
Co-fractionation
Co-localization
Co-purification
FRET
PCA
Two-hybrid
Biochemical_activity
Co-crystal_structure
Far_western
Protein_peptide
Protein_RNA
Reconstituted_complex
Y1H
Directed_Y1H Text
Protein_DNA
Throughput UNIQUE High_throughput //Need to define in context of physical interactions
Low_throughput //Same as above
Library_info Library UNIQUE ?Library //This could also just be ?Text. Doesn't look like ?Library class is used.
Origin From_laboratory UNIQUE ?Laboratory //XREF by making a Reagents tag in the ?Laboratory model?
From_company UNIQUE ?Text //We don't currently have a ?Company class. Should we?
Confidence ?Text //This can accommodate the great variety of language used to expressed this, if curated.
Paper ?Paper XREF Interaction //Should this XREF also be updated to Physical_interaction?
Remark ?Text #Evidence
Physical Interaction Model v1
This version of the model treats each instance of a physical interaction as a separate entity.
?Physical_interaction Evidence #Evidence
Interactor Bait PCR_bait UNIQUE ?PCR_product XREF Interaction_bait //Change XREF tag to Physical...
Sequence_bait UNIQUE ?Sequence XREF Interaction_bait
Bait_overlapping_CDS ?CDS XREF Interaction_bait #Evidence
Bait_overlapping_gene ?Gene XREF Interaction_bait #Evidence
Target PCR_target UNIQUE ?PCR_product XREF Interaction_target //Change Target to Hit? Also change XREF as above?
Sequence_target UNIQUE ?Sequence XREF Interaction_target
Target_overlapping_CDS ?CDS XREF Interaction_target #Evidence
Target_overlapping_gene ?Gene XREF Interaction_target #Evidence
Experiment_type Affinity_capture-luminescence
Affinity_capture-MS
Affinity_capture-RNA
Affinity_capture-Western
Co-fractionation
Co-localization
Co-purification
FRET
PCA
Two-hybrid
Biochemical_activity
Co-crystal_structure
Far_western
Protein_peptide
Protein_RNA
Reconstituted_complex
Y1H
Directed_Y1H Text
Protein_DNA
Throughput UNIQUE High_throughput //Need to define in context of physical interactions
Low_throughput //Same as above
Library_info Library UNIQUE ?Library //This could also just be ?Text. Doesn't look like ?Library class is used.
Origin Species UNIQUE ?Species
From_laboratory UNIQUE ?Laboratory //XREF by making a Reagents tag in the ?Laboratory model?
From_company UNIQUE ?Text //We don't currently have a ?Company class. Should we?
Confidence Confidence_level UNIQUE Float //Do we need this in the ?Physical_interaction model?
P_value UNIQUE Float //Same as above.
Log_likelihood_score UNIQUE Float //Same as above.
Interaction_frequency UNIQUE Int //This would hold the Int data in the existing Library_screened tag.
Interactome_type UNIQUE Interactome_core_1 //As defined in Li et al., 2004
Interactome_core_2
Interactome_core_3
Paper ?Paper XREF Interaction //Should this XREF also be updated to Physical_interaction?
Remark ?Text #Evidence
Physical Interaction Model v2
This version of the model gives a single interaction ID to two interacting entities, but each instance, or evidence for the interaction, is added in the #Interaction_info under the corresponding Experiment_type.
?Physical_interaction Evidence #Evidence
Interactor Bait PCR_bait UNIQUE ?PCR_product XREF Interaction_bait //Change XREF tag to Physical...
Sequence_bait UNIQUE ?Sequence XREF Interaction_bait
Bait_overlapping_CDS ?CDS XREF Interaction_bait #Evidence
Bait_overlapping_gene ?Gene XREF Interaction_bait #Evidence
Target PCR_target UNIQUE ?PCR_product XREF Interaction_target //Change Target to Hit? Also change XREF as above?
Sequence_target UNIQUE ?Sequence XREF Interaction_target
Target_overlapping_CDS ?CDS XREF Interaction_target #Evidence
Target_overlapping_gene ?Gene XREF Interaction_target #Evidence
Experiment_type Affinity_capture-luminescence #Interaction_info
Affinity_capture-MS #Interaction_info
Affinity_capture-RNA #Interaction_info
Affinity_capture-Western #Interaction_info
Co-fractionation #Interaction_info
Co-localization #Interaction_info
Co-purification #Interaction_info
FRET #Interaction_info
PCA #Interaction_info
Two-hybrid #Interaction_info
Biochemical_activity #Interaction_info
Co-crystal_structure #Interaction_info
Far_western #Interaction_info
Protein_peptide #Interaction_info
Protein_RNA #Interaction_info
Reconstituted_complex #Interaction_info
Y1H #Interaction_info
Directed_Y1H Text #Interaction_info
Protein_DNA #Interaction_info
Remark ?Text #Evidence
#Interaction_info Interaction_RNAi ?RNAi XREF Interaction
Effector ?Gene //master, upstream
Effected ?Gene //subject, downstream
Non_directional ?Gene //e.g. synthetic interactions - Igor
Interaction_phenotype ?Phenotype XREF Interaction
Throughput UNIQUE High_throughput
Low_throughput
Library_info Library UNIQUE ?Library
Origin Species UNIQUE ?Species
From_laboratory UNIQUE ?Laboratory
From_company ?Text
Confidence Confidence_level UNIQUE Float
P_value UNIQUE Float
Log_likelihood UNIQUE Float
Interaction_frequency UNIQUE Int
Interactome_type UNIQUE Interactome_core_1
Interactome_core_2
Interactome_noncore
Paper ?Paper XREF Interaction
Physical Interaction Model v3
This model keeps the physical interaction as part of the general ?Interaction model with the details again going into the #Interaction_info. The #Interaction_info would now contain the information about bait/hit directionality.
?Interaction Evidence #Evidence
Interactor ?Gene XREF Interaction #Interactor_info
Interaction_type Genetic #Interaction_info
Regulatory #Interaction_info
No_interaction #Interaction_info
Predicted_interaction #Interaction_info
Physical_interaction #Interaction_info
Suppression #Interaction_info
Enhancement #Interaction_info
Synthetic #Interaction_info
Epistasis #Interaction_info
Mutual_enhancement #Interaction_info
Mutual_suppression #Interaction_info
DB_info Database ?Database ?Database_field ?Accession_number
Remark ?Text #Evidence
#Interaction_info Interaction_RNAi ?RNAi XREF Interaction
Effector ?Gene //master, upstream
Effected ?Gene //subject, downstream
Bait PCR_bait UNIQUE ?PCR_product XREF Interaction_bait //Change XREF tag to Physical...
Sequence_bait UNIQUE ?Sequence XREF Interaction_bait
Bait_overlapping_CDS ?CDS XREF Interaction_bait #Evidence
Bait_overlapping_gene ?Gene XREF Interaction_bait #Evidence
Target PCR_target UNIQUE ?PCR_product XREF Interaction_target //Change Target to Hit? Also change XREF as above?
Sequence_target UNIQUE ?Sequence XREF Interaction_target
Target_overlapping_CDS ?CDS XREF Interaction_target #Evidence
Target_overlapping_gene ?Gene XREF Interaction_target #Evidence
Experiment_type Affinity_capture-luminescence
Affinity_capture-MS
Affinity_capture-RNA
Affinity_capture-Western
Co-fractionation
Co-localization
Co-purification
FRET
PCA
Two-hybrid
Biochemical_activity
Co-crystal_structure
Far_western
Protein_peptide
Protein_RNA
Reconstituted_complex
Y1H
Directed_Y1H Text
Protein_DNA
Throughput UNIQUE High_throughput
Low_throughput
Library_info Library UNIQUE ?Library //This could also just be ?Text. Doesn't look like ?Library class is used.
Origin Species UNIQUE ?Species
From_laboratory UNIQUE ?Laboratory //XREF by making a Reagents tag in the ?Laboratory model?
From_company UNIQUE ?Text //We don't currently have a ?Company class. Should we?
Confidence Confidence_level UNIQUE Float
P_value UNIQUE Float
Log_likelihood_score UNIQUE Float
Interaction_frequency UNIQUE Int //This would hold the Int data in the existing Library_screened tag.
Interactome_type UNIQUE Interactome_core_1 //As defined in Li et al., 2004
Interactome_core_2
Interactome_core_3
Paper ?Paper XREF Interaction //Should this XREF also be updated to Physical_interaction?
Remark ?Text #Evidence
OA interface
- Tab 1
- PGID Same in new OA
- Interaction ID
- A new interaction ID is generated by clicking on 'new'. when 'duplicate', the ID from old entry will be in the field, but need to be deleted in order to get an new ID.
- Interaction ID will be assigned by cronjob daily at 4 am. This is for curators who would like to use 'Duplicate' to generate new objects with similar field entries and erase the Interaction IDs to let the cron job add new IDs overnight. The cronjob is at /home/acedb/xiaodong/assigning_interaction_ids/assign_interaction_ids.pl and we need to change it based on the new table structure -- J
- interaction field autocompletes now key off of the int_index table used by the interaction_ticket.cgi, it no longer keys off of the int_name table from this field in the interaction OA, which makes the rest of this line obsolete. (OBSOLETE If you mistakenly make a typo and assign a correct ID's value to some other ID, you will _not_ be able to bring it back (because it's an ontology) without going to postgres directly and editing the int_name and int_name_hst tables by pgid (in postgres called joinkey). You'll have to note the pgid and then manually change it in postgres. end obsolete)
- Non_directional In new OA, there will be a separate field for each interactor type: Gene, Sequence, CDS, PCR_Product, or Protein. Since "old" interaction objects only contain genes, any genes that are part of Interaction obejcts in which the Non_directional toggle was activated will need to move to the "Non-directional Gene(s)" field. if toggle is on, move all effector + effected genes to the new non-directional gene field. get rid of this field in new OA
- toggle OFF (default), means interaction is directional. there is effected/effector parties involve in the object.
- toggle ON (color change to red by click) means interaction is non_directional
- Interaction Type//dropdown list with 11 types showing in .ace template
- In the new OA, these 11 types will be mapped as follows:
- "Genetic" will become "Genetic - Genetic Interaction"
- "Regulatory" will remain "Regulatory"
- "No_interaction" will become "Genetic - No_interaction"
- "Predicted_interaction" will become "Predicted"
- "Physical_interaction" will become "Physical"
- "Suppression" will become "Genetic - Suppression"
- "Enhancement" will become "Genetic - Enhancement"
- "Synthetic" will become "Genetic - Synthetic"
- "Epistasis" will become "Genetic - Epistasis"
- "Mutual_enhancement" will become "Genetic - Mutual_enhancement"
- "Mutual_suppression" will become "Genetic - Mutual_suppression"
- Effected Gene //autocomplete WBGene, multiontology, corresponding to interactor in .ace file. order does not matter when dump to interactors
- In new OA, this field will be called "Affected Gene(s)" -- C If non-directional is on, move to non-directional gene, otherwise leave here
- Effected Variation //WBGene, WBVar, multiontology, autocomplete on Variation, store in separate lines ->.ace, Interactor "WBGene" Variation "WBVar"
- In new OA, this field will be called "Affected Variation(s)"; the dumping script will attempt to map these variations to their respetive genes and place the variation in that gene's interactor_info hash -- C
- In new OA, the gene(s) affiliated with this variation will become the "Affected Gene(s)" for this interaction.
- Effected Transgene_Name //ontology, autocomplete transgene object name, eg iaIs3.
- In new OA, all transgenes will go into the Transgene(s) field. merge both effector + effected.
- Effected Transgene_Gene // multi-ontology, autocomplete WBGene->.ace, Interactor "WBGene" Transgene "id". In case of multi genes, WBGene is followed by same transgene id. One wbgene for each .ace line ? Make sure you really want it this way, we can go with product/promoter if that's what you want, just make sure it's what you want. It matters having extra fields and scrolling and so forth. You'll see when the text fields become multi-ontology and ontology.
- In new OA, all transgenes will automatically be mapped to their associated genes based on all genes in the transgene OA's gene + driven_by_gene field. remove this field.
- In new OA, genes from the "Effected Transgene Gene" field will move to the "Affected Gene" field.
- Effected Other Type //dropdown list of 'Chemicals' and 'Transgene'
- In new OA, these will be named "Affected Other Type" and dump as remark.
- Effected Other //free text field now, however, when entering chemicals make sure to enter common names followed by mesh IDs in parenthesis for later ontologinization.
- In new OA, these will be named "Affected Other" and dump as remark.
- Effector Gene //autocomplete WBGene, multiontology. corresponding to interactor in .ace file. order does not matter when dump to interactors
- In new OA, these genes will all go in the "Effector Gene(s)" field. If non-directional is on, move to non-directional gene, otherwise leave here
- Effector Variation //WBGene, WBVar, autocomplete multiontology on variation, store in separate lines ->.ace, Interactor "WBGene" Variation "WBVar". use name server to map variation to gene, or the file Karen gave you to map variation to gene for variation OA.
- In new OA, this field will be the same; the dumping script will attempt to map these variations to their respetive genes and place the variation in that gene's interactor_info hash -- C
- In new OA, the gene(s) affiliated with this variation will become the "Effector Gene(s)" for this interaction.
- Effector Transgene_Name //autocomplete name, ontology
- In new OA, all transgenes will go into the Transgene(s) field. merge both effector + effected.
- Effector Transgene_Gene // autocomplete WBGene, multi-ontology, ->.ace, Interactor "WBGene" Transgene "id". In case of multi genes, WBGene is followed by same transgene id.
- In new OA, all transgenes will automatically be mapped to their associated genes based on all genes in the transgene OA's gene + driven_by_gene field. remove this field.
- In new OA, genes from the "Effector Transgene Gene" field will move to the "Effector Gene" field.
- Effector Other Type //dropdown list of 'Chemicals' and 'Transgene'
- In new OA, these will stay the same and dump as Remark
- Effector Other //free text field
- two fields above will be dumped in remark field.
- In new OA, these will stay the same and dump as Remark
- Effected Gene //autocomplete WBGene, multiontology, corresponding to interactor in .ace file. order does not matter when dump to interactors
Note: Gene, Variation, and Transgene_Gene all refer to different genes. There is no pairing problem.
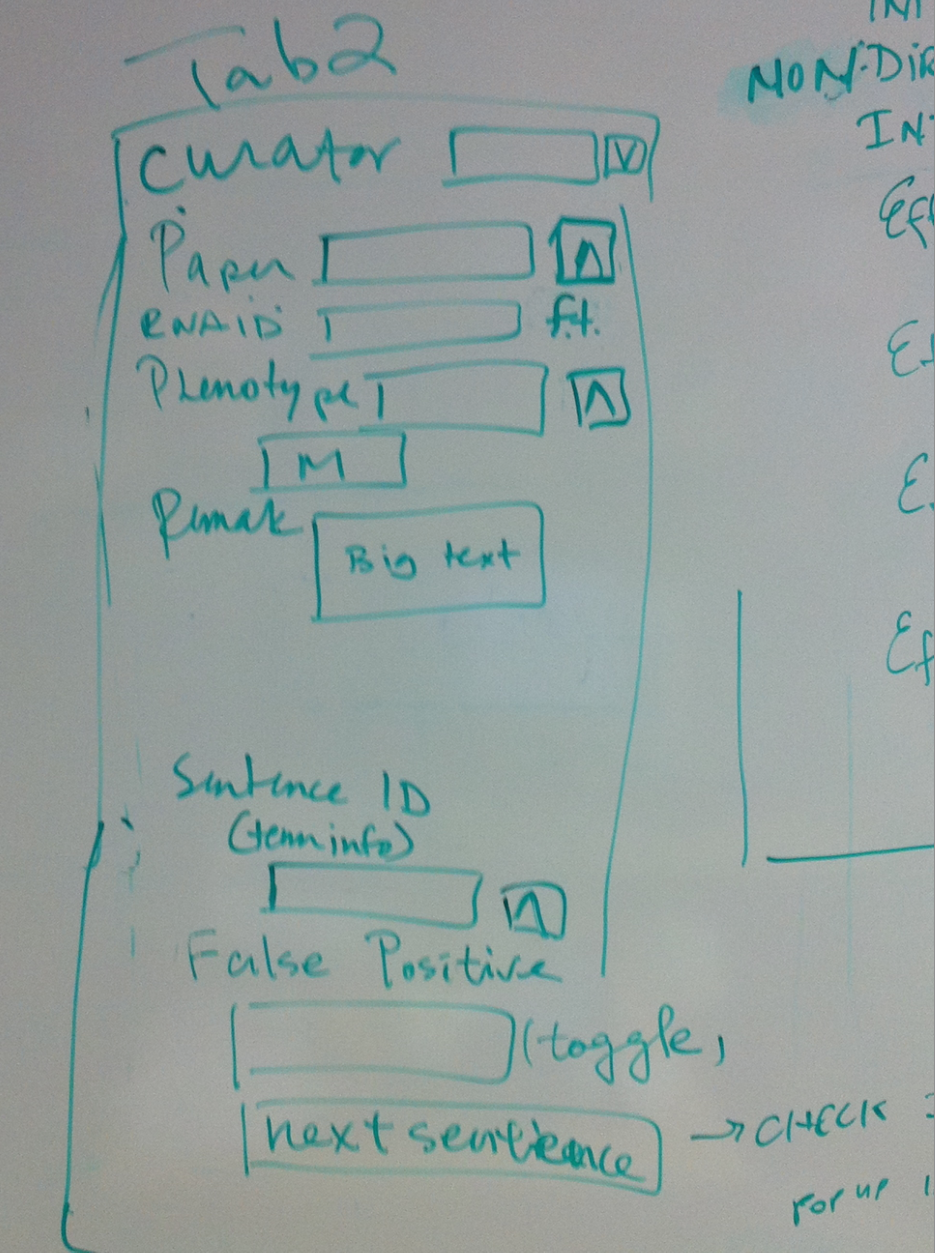
- Tab 2
- Curator//dropdown list Same in new OA
- Paper//ontology Same in new OA
- RNAi ID//free text field In new OA, this RNAi object will fill the "RNAi" field if it is not from WBPaper00029258 (hence large scale); if it is from WBPaper00029258, this will go in the "Large scale RNAi" field as free text
- Phenotype//multiontology In new OA, this Phenotype object will fill the "Interaction phenotype" field. keep OA field the same, change dumper.
- Remark//big text Same in new OA
- Sentence ID//sentence shows in term info Same in new OA
- False Positive//toggle, will not give an id or no dump if the sentence is false positive, containing no interaction info Same in new OA
Old Table Names
'Non_directional' = int_nondirectional
'Effector Transgene Name' = int_transgeneone
'Effector Transgene Gene' = int_transgeneonegene
'Affected Transgene Name' = int_transgenetwo
'Affected Transgene Gene' = int_transgenetwogene
'Deviation from expectation' = int_deviation
'Neutrality function' = int_neutralityfxn
'Intragenic Effector Variation(s)' = int_intravariationone
'Intragenic Affected Variation(s)' = int_intravariationtwo
'Effector Other Type' = int_otheronetype
'Affected Other Type' = int_othertwotype
Removed OA fields
- As of 2-19-2015
TAB3
- Intragenic Effector Variation(s) - ?Variation (MultiOntology) int_intravariationone
- Genes for this field need to be mapped to a gene at the dump stage; Genes that map to the variation will be dumped as the "Interactor_overlapping_gene" as follows:
- Dumps as: Interactor_overlapping_gene <Mapped Gene> Intragenic_effector_variation <Variation>
- For Variations that don't map to a gene in the interaction, Dump as: Unaffiliated_variation <Variation>
- Intragenic Affected Variation(s) - ?Variation (MultiOntology) int_intravariationtwo
- Genes for this field need to be mapped to a gene at the dump stage; Genes that map to the variation will be dumped as the "Interactor_overlapping_gene" as follows:
- Dumps as: Interactor_overlapping_gene <Mapped Gene> Intragenic_affected_variation <Variation>
- For Variations that don't map to a gene in the interaction, Dump as: Unaffiliated_variation <Variation>
TAB4
- Deviation from expectation - Big text int_deviation Dumps as: Deviation_from_expectation <Big_Text>
- Neutrality function - (Dropdown) int_neutralityfxn options are:
- Multiplicative Dumps as: Multiplicative
- Additive Dumps as: Additive
- Minimal Dumps as: Minimal
- Effector Other Type - (Dropdown) int_otheronetype options are: Chemical or Transgene, int_otheronetype Dumps as (see next line)
- Affected Other Type - (Dropdown) int_othertwotype options are: Chemical or Transgene, int_othertwotype Dumps as (see next line)
How to use:
Deviation from expectation is a free-text field where, at the curator's discretion, a curator can describe why a genetic interaction is an interaction, i.e. why it is an unexpected result warranting an interaction. This could be as simple as "The life spans were synergistic" or "Neither mutation alone exhibited a phenotype".
Neutrality function is closely related to the "Deviation from expectation" field, where the curator can decide, if applicable, which "Neutrality function" applies to this genetic interaction. The choices in this drop down field are "Multiplicative", "Additive", and "Minimal". "Multiplicative" means that the authors expected to see a quantifiable phenotype in the double mutant that was the mathematical product of the quantified phenotypes of the individual mutations. For example, one mutant extends life span by 20%, and another extends it by 30%. With a multiplicative neutrality function, we would expect the double mutant to have (1.2 * 1.3 = 1.56) about 56% extended life span. Alternatively, in the "Additive" neutrality function, the authors might expect the sum of the effects (A + B - 1) or (1.2 + 1.3 - 1 = 1.5) or a 50% extended life span. The "Minimal" neutrality function assumes that the double mutant will be as severe as the most severe single mutant, in this case (1.3) or 30% extended. Therefore, any life span extension beyond 30%, for example, in the double mutant would be considered "surprising" and therefore, an interaction.
Effector/Affected Other Type and Effector/Affected Other fields: these fields allow for the curation of Chemicals, Transgenes, or other entities that don't exists as proper WormBase/ACEDB objects, for example transgenes that only express human proteins. The "Other Type" fields allow for the selection of "Chemical" or "Transgene", the identity of which would go into the "Other" fields. This, ideally, will get phased out as chemicals are generated in the Molecule OA and transgenes in the Transgene OA, thereby allowing them to be entered into ontology-based "Transgene" or "Molecule" fields.
.ace template: (OUTDATED)
- Interaction : ""
- Interactor "WBGene" Variation ""
- Interactor "WBGene" Transgene ""
- Interactor "WBGene"
- Interaction_type Genetic Effector ""
- Interaction_type Genetic Effected ""
- Interaction_type Genetic Non_directional ""
- Interaction_type Genetic Interaction_RNAi ""
- Interaction_type Genetic Interaction_phenotype ""
- Interaction_type Regulatory Effector ""
- Interaction_type Regulatory Effected ""
- Interaction_type Regulatory Non_directional ""
- Interaction_type Regulatory Interaction_RNAi ""
- Interaction_type Regulatory Interaction_phenotype ""
- Interaction_type No_interaction Effector ""
- Interaction_type No_interaction Effected ""
- Interaction_type No_interaction Non_directional ""
- Interaction_type No_interaction Interaction_RNAi ""
- Interaction_type No_interaction Interaction_phenotype ""
- Interaction_type Predicted_interaction Non_directional ""
- Interaction_type Predicted_interaction Interaction_RNAi ""
- Interaction_type Predicted_interaction Interaction_phenotype ""
- Interaction_type Physical_interaction Effector ""
- Interaction_type Physical_interaction Effected ""
- Interaction_type Physical_interaction Interaction_RNAi ""
- Interaction_type Physical_interaction Interaction_phenotype ""
- Interaction_type Suppression Effector ""
- Interaction_type Suppression Effected ""
- Interaction_type Suppression Interaction_RNAi ""
- Interaction_type Suppression Interaction_phenotype ""
- Interaction_type Enhancement Effector ""
- Interaction_type Enhancement Effected ""
- Interaction_type Enhancement Interaction_RNAi ""
- Interaction_type Enhancement Interaction_phenotype ""
- Interaction_type Synthetic Non_directional ""
- Interaction_type Synthetic Interaction_RNAi ""
- Interaction_type Synthetic Interaction_phenotype ""
- Interaction_type Epistasis Effector ""
- Interaction_type Epistasis Effected ""
- Interaction_type Epistasis Interaction_RNAi ""
- Interaction_type Epistasis Interaction_phenotype ""
- Interaction_type Mutual_enhancement Non_directional ""
- Interaction_type Mutual_enhancement Interaction_RNAi ""
- Interaction_type Mutual_enhancement Interaction_phenotype ""
- Interaction_type Mutual_suppression Non_directional ""
- Interaction_type Mutual_suppression Interaction_RNAi ""
- Interaction_type Mutual_suppression Interaction_phenotype ""
- Paper ""
- Remark ""
interaction objects source file
- there are 9242 interaction objects dumped from WS220 on Monday, 10/01/2010
- Juancarlos's parse results from this file:
/home/postgres/work/pgpopulation/interaction/20101004_xiaodong_start/out
There are two interactions in postgres, but not the .ace file : In postgres, no ace WBInteraction0008637 In postgres, no ace WBInteraction0008638//will be OA
There are 1290 interactions in .ace file not in postgres (so I imagine these are what we should read in ?)//these are RNAi based interaction objects, we want to include them in OA
There are >40000 interactions that have a ticket and are in neither .ace nor postgres.//these 398,619 interactions are from two large scale papers
Also there are interaction data in postgres without an interaction ID//will be assigned id from WBInteraction0500001.
Some Notes for gene_gene_interaction
Outdated notes on large scale interaction files
ready_to_upload large scale data file: Large_scale_interactions_WS231.ace is located on tazendra: /home/acedb/xiaodong/oa_interactions_dumper should be uploaded for every uploadlarge scale .ace file needs to be checkd again dead genes for each upload using scripts in the same directory: find_invalid_genes_largescale.pl* - 08/31/2012 for upload WS234- before run, change source file (in file) to latest large scale .ace version (in scripts)
- dead/merged genes will appear on screen. replace merged genes, leave dead/dead genes for now
- change large scale.ace file name to latest upload version
then upload
Citace upload notes
- 2011.1.27
- caught-up at acedb reading: WBInteraction0500069 (Karen), new line in remark field. fixed in .ace file.
- some confusion on ids. found out ticket issuer was using sandbox data. fixed.
- 2011.5.4
- Karen needed to update variation_wbgene file on tazendra: /home/acedb/jolene/WS_AQL_queries/Variation_gene.txt
- Juancarlos changed the dumper to ignore line breaks and double spaces in remark field for dumping ace file