Difference between revisions of "Contributing Phenotype Connections"
m |
|||
| (18 intermediate revisions by the same user not shown) | |||
| Line 1: | Line 1: | ||
The following documentation explains the use and backend for the Phenotype form found here: | The following documentation explains the use and backend for the Phenotype form found here: | ||
| − | + | https://wormbase.org//submissions/phenotype.cgi | |
| Line 70: | Line 70: | ||
=== PubMed ID === | === PubMed ID === | ||
| − | This form requires the entry of a PubMed ID (or D.O.I. if no PubMed ID exists) for the publication in which the phenotype data was originally reported. PubMed IDs may be entered manually with or without the "pmid" prefix, e.g. "pmid6616618" or simply "6616618". Once an ID has been entered and recognized, a term information box will appear in the upper right-hand corner of the screen with additional information about the paper including title, journal, year, and | + | This form requires the entry of a PubMed ID (or D.O.I. if no PubMed ID exists) for the publication in which the phenotype data was originally reported. PubMed IDs may be entered manually with or without the "pmid" prefix, e.g. "pmid6616618" or simply "6616618". Once an ID has been entered and recognized, a term information box will appear in the upper right-hand corner of the screen with additional information about the paper including title, journal, year, and last name of the first author: |
| − | [[File: | + | [[File:PubMed_ID_2881214_term_info_Apr_20_2018.png|500px]] |
| Line 134: | Line 134: | ||
| − | If a recognized gene name is selected from the autocomplete suggestions, a term information box will appear in the upper right corner of the screen displaying basic information about the gene with a link to the respective WormBase gene page: | + | If a recognized gene name is selected from the autocomplete suggestions, a term information box will appear in the upper right corner of the screen displaying basic information about the gene with a link (locus name is hyperlinked) to the respective WormBase gene page: |
| Line 140: | Line 140: | ||
| − | + | Once a gene has been selected (or typed in), an "RNAi reagent" field and a "Species" field will appear: | |
| − | [[File: | + | [[File:Phenotype_form_RNAi_reagent_and_species_fields_Feb_2018.png|600px]] |
| − | Because RNA interference experiments can now be carried out in several species, the form has a species selection to specify which species the RNAi experiment(s) was/were carried out in, with a default to Caenorhabditis elegans. | + | You may enter a library name and gene name (e.g. "Ahringer dpl-1 clone"), the commercial clone well location (e.g. "Source BioScience II-6H13") or the sequence of the RNAi clone (e.g. "ACGTATGCTAG...."). Because RNA interference experiments can now be carried out in several species, the form has a species selection to specify which species the RNAi experiment(s) was/were carried out in, with a default to Caenorhabditis elegans. Note that as soon as an entry is made in the "RNAi target gene" field, an additional "RNAi target gene" field will appear, allowing submission of multiple RNAi target genes simultaneously. If an entry is made in the second RNAi target gene field (or more than one genetic perturbation is entered in general), a notice will appear asking the submitter to indicate if the multiple genetic perturbations represent part of a complex genotype (single phenotype experiment) or multiple independent phenotype experiments (each with the indicated phenotype); see below. |
==== Allele ==== | ==== Allele ==== | ||
| Line 167: | Line 167: | ||
| − | [[File: | + | [[File:Phenotype_form_unrecognized_allele_Feb_2018.png|600px]] |
| − | If you are entering a "new" allele name, click anywhere outside the field to set the name, and continue with your submission. An allele curator will be notified and may contact you to confirm the existence of this allele. | + | If you are entering a "new" allele name, click anywhere outside the field to set the name, and continue with your submission. An allele curator will be notified and may contact you to confirm the existence of this allele. Note that as soon as an entry is made in the "Allele" field, an additional "Allele" field will appear, allowing submission of multiple alleles simultaneously. If an entry is made in the second Allele field (or more than one genetic perturbation is entered in general), a notice will appear asking the submitter to indicate if the multiple genetic perturbations represent part of a complex genotype (single phenotype experiment) or multiple independent phenotype experiments (each with the indicated phenotype); see below. |
==== Transgene ==== | ==== Transgene ==== | ||
| Line 195: | Line 195: | ||
| + | The "Gene Causing Phenotype" field has an autocomplete feature similar to the "RNAi target gene" field, allowing recognized gene names as well as unrecognized (new) gene names with a notice similar to that of the "RNAi target gene" field. A notice will appear to the right of the field to indicate if a transgene name that has been entered is not recognized by WormBase: | ||
| − | A | + | |
| + | [[File:Phenotype_form_unrecognized_transgene_notice_1-20-2016.png|600px]] | ||
| + | |||
| + | |||
| + | If you are entering a "new" transgene name, click anywhere outside the field to set the name, and continue with your submission. A curator will be notified and may contact you to confirm the existence of this transgene. Note that as soon as an entry is made in the "Transgene" field, an additional "Transgene" field will appear, allowing submission of multiple transgenes simultaneously. If an entry is made in the second Transgene field (or more than one genetic perturbation is entered in general), a notice will appear asking the submitter to indicate if the multiple genetic perturbations represent part of a complex genotype (single phenotype experiment) or multiple independent phenotype experiments (each with the indicated phenotype); see below. | ||
| − | [[File: | + | ==== Entering multiple genetic perturbations ==== |
| + | |||
| + | For convenience, the phenotype form allows data submitters to submit multiple genetic perturbations simultaneously in one phenotype data submission. This way, contributors can indicate, in one submission, that multiple experiments each with distinct alleles or RNAi knockdowns, for example, each resulted in the observed phenotype(s) compared to control. Alternatively, the intention may be to indicate that multiple genetic perturbations were simultaneously introduced into some genetic background and a phenotype was observed for a double mutant, for example. To distinguish between these two interpretations, as soon as two or more genetic perturbations are entered into the form, a notice appears asking the user to specify which of these two situations applies for the multiple genetic perturbations entered: | ||
| + | |||
| + | |||
| + | [[File:Phenotype_form_multiple_perturbations_Feb_2018.png|600px]] | ||
| + | |||
| + | If each genetic perturbation entered represents an individual, independent phenotype experiment, each resulting in the observed phenotype(s), select (click on) the radio button to the left of "Multiple individual experiments", so that the annotation can be properly entered into the WormBase database. On the other hand, if the multiple genetic perturbations entered represents a single complex genotype experiment (e.g. a double or triple mutant phenotype) in which all perturbations were simultaneously assessed for a phenotype in a single organism or isogenic population, select (click on) the radio button to the left of "Single complex genotype experiment". | ||
| − | + | Note that entering multiple perturbations in a single submission will prevent the form from accepting entries in the "Inheritance Pattern" or "Mutation Effect" fields in the Optional section (see below). | |
=== Phenotype(s) === | === Phenotype(s) === | ||
| Line 294: | Line 306: | ||
| − | [[File: | + | [[File:Phenotype_form_submission_preview_Feb_2018.png|700px]] |
| − | |||
=== Submission === | === Submission === | ||
| Line 302: | Line 313: | ||
| − | [[File: | + | [[File:Phenotype_form_submission_confirmation_Feb_2018.png|700px]] |
| Line 308: | Line 319: | ||
| − | [[File: | + | [[File:Phenotype_form_submission_confirmation_email_Feb_2018.png|700px]] |
| Line 314: | Line 325: | ||
| − | [[File: | + | [[File:Phenotype_form_submission_retraction_confirmation_Feb_2018.png|500px]] |
| − | |||
= WormBase Internal Documentation = | = WormBase Internal Documentation = | ||
| Line 360: | Line 370: | ||
* Your E-mail Address --> Community Curator Email (app_communitycuratoremail) | * Your E-mail Address --> Community Curator Email (app_communitycuratoremail) | ||
* PubMed ID --> Unregistered Paper (if unrecognized) (app_unregpaper) | * PubMed ID --> Unregistered Paper (if unrecognized) (app_unregpaper) | ||
| − | * | + | * Allele --> Unregistered Variation (if unrecognized or entire genotype if multiple perturbations entered for a single complex genotype experiment) (app_unregvariation) |
* Transgene --> Unregistered Transgene (if unrecognized) (app_unregtransgene) | * Transgene --> Unregistered Transgene (if unrecognized) (app_unregtransgene) | ||
* "NO DUMP" (app_nodump) and "Needs Review" (app_needsreview) toggled ON by default for every submission | * "NO DUMP" (app_nodump) and "Needs Review" (app_needsreview) toggled ON by default for every submission | ||
| Line 378: | Line 388: | ||
* N/A --> Curator (rna_curator) defaults to "Community Curator (WBPerson29819)" for every submission | * N/A --> Curator (rna_curator) defaults to "Community Curator (WBPerson29819)" for every submission | ||
* PubMed ID --> Paper (as WBPaper ID) (rna_paper) | * PubMed ID --> Paper (as WBPaper ID) (rna_paper) | ||
| − | * RNAi | + | * RNAi target gene --> Remark (rna_remark) as "<WBGene ID>" |
| + | ** RNAi reagent --> DNA text (rna_dnatext) | ||
| + | ** RNAi species --> Species (rna_species) | ||
* Control Strain (Optional section) --> Strain (rna_strain) | * Control Strain (Optional section) --> Strain (rna_strain) | ||
* Control Genotype (Optional section) --> Genotype (rna_genotype) | * Control Genotype (Optional section) --> Genotype (rna_genotype) | ||
| Line 397: | Line 409: | ||
| − | Currently (as of | + | Currently (as of February 2018), all submissions through the Phenotype form will send an e-mail to the submitter and to Chris Grove and Gary Schindelman and the "NO DUMP" and "Needs Review" toggles will be turned ON by default in the respective OA's. Community curation submissions through this form can be queried by querying for the "Community Curator (WBPerson29819)" and those in need of review can be queried via the "Needs Review" toggle. A history of all submissions to the form can be viewed at: |
http://tazendra.caltech.edu/~azurebrd/cgi-bin/data/phenotype_history.html | http://tazendra.caltech.edu/~azurebrd/cgi-bin/data/phenotype_history.html | ||
Latest revision as of 21:32, 8 April 2020
The following documentation explains the use and backend for the Phenotype form found here:
https://wormbase.org//submissions/phenotype.cgi
Contents
User Guide
Finding the Phenotype form
To reach the Phenotype form from the WormBase homepage, click on the "Submit Data" link below the upper right search bar:
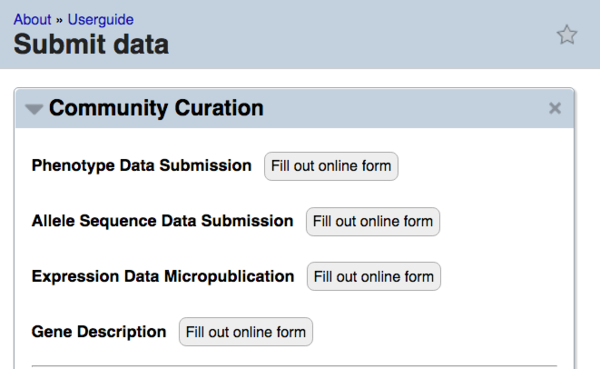
This link leads you to the "Submit Data" page. On this page, the "Community Curation" widget has a link to the Phenotype form via the "Fill out online form" button to the right of "Phenotype Data Submission":
Overview
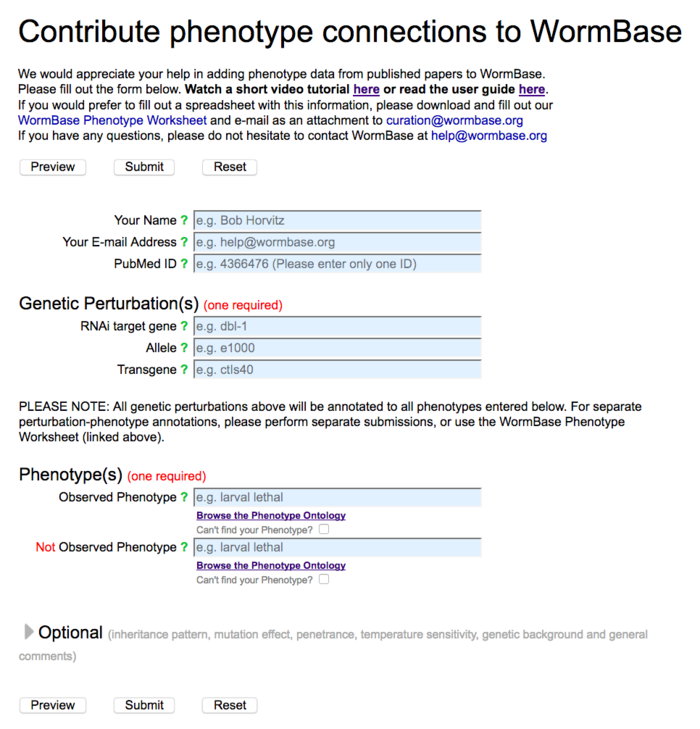
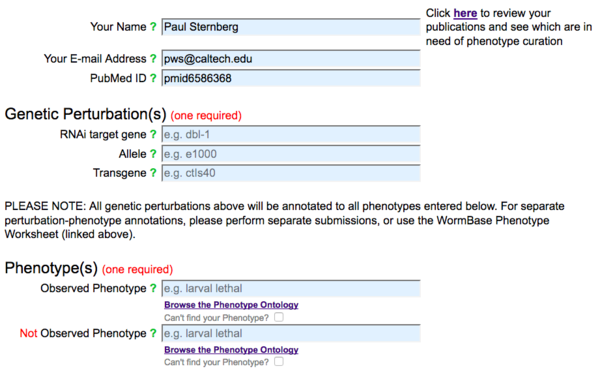
The first part of the Phenotype form carries all of the fields that are required for data submission: a submitter's name and e-mail address, a publication's PubMed ID, a genetic perturbation (at least one of RNAi, allele or transgene), and a phenotype (observed or not observed):
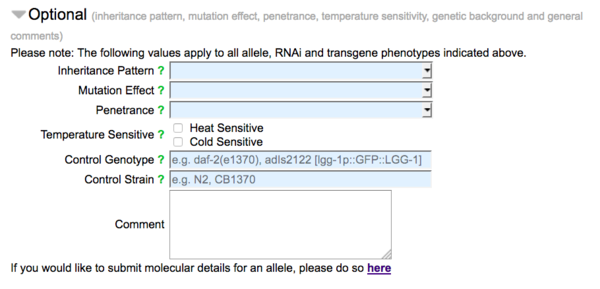
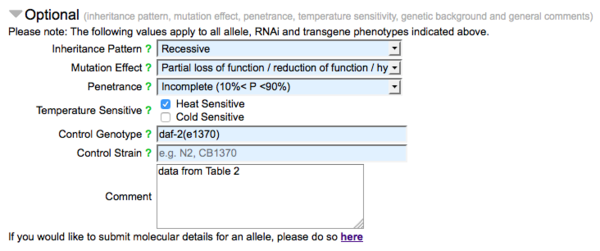
The second part of the form (made visible by clicking on the header "Optional") carries fields for optional data, pertaining to the nature of the allele, RNAi clone, or transgene with respect to the observed phenotypes indicated in the form:
A drop down list of options for inheritance pattern (e.g. dominant, recessive), mutation effect (e.g. loss of function, null, gain of function) and penetrance (roughly categorized as low, incomplete, high, and complete) are available. Check boxes are also available to indicate the temperature sensitivity of the allele. There are also fields for genetic background (genotype or strain) and general comments for the submission. The bottom of the "Optional" section has a link to the WormBase Allele-Sequence form, by which sequence details of alleles may be reported.
Required information
The form minimally requires a submitter's name and e-mail address, a publication's PubMed ID, a genetic perturbation (allele, RNAi, or transgene), and a phenotype (observed or NOT observed).
Your Name & E-mail Address
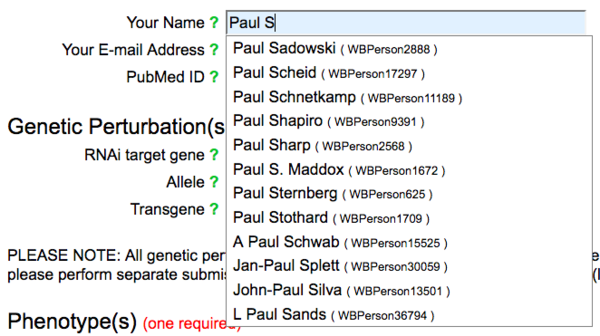
The name field has an autocomplete functionality, requiring the selection of a name from the list of registered WormBase Persons.
If your name is not registered with WormBase, click on the green question mark to the left of the name field:
to open up the help text in the term information box in the upper right-hand corner of the screen and click on the "Person Update Form" link to register.
Once a name has been selected, a term information box will appear in the upper right-hand corner of the page with additional information about this WormBase Person:
including a link to their WormBase Person page, as well as a link to the WBPerson publication summary page (see below). If WormBase has your e-mail address on file, it will automatically be filled into the e-mail field, otherwise you may add (or correct) your e-mail address manually.
PubMed ID
This form requires the entry of a PubMed ID (or D.O.I. if no PubMed ID exists) for the publication in which the phenotype data was originally reported. PubMed IDs may be entered manually with or without the "pmid" prefix, e.g. "pmid6616618" or simply "6616618". Once an ID has been entered and recognized, a term information box will appear in the upper right-hand corner of the screen with additional information about the paper including title, journal, year, and last name of the first author:
To assist in the look-up of PubMed IDs relevant to you, once your name has been selected a notice will appear to the right of the name field with a link to a summary of your publications and their associated PubMed IDs:
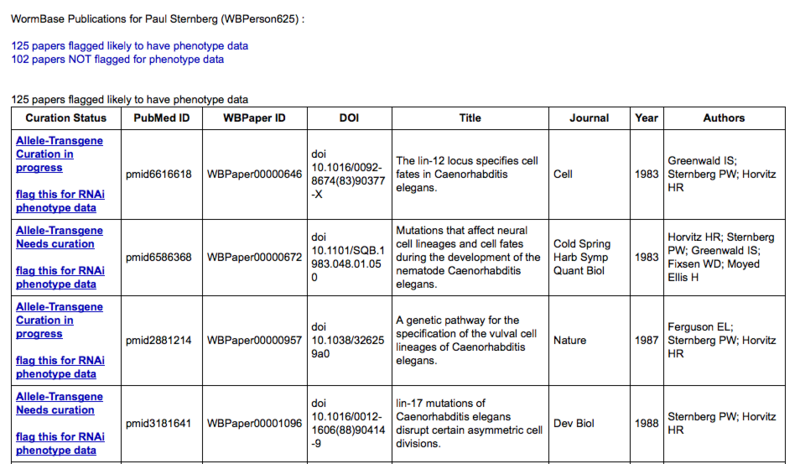
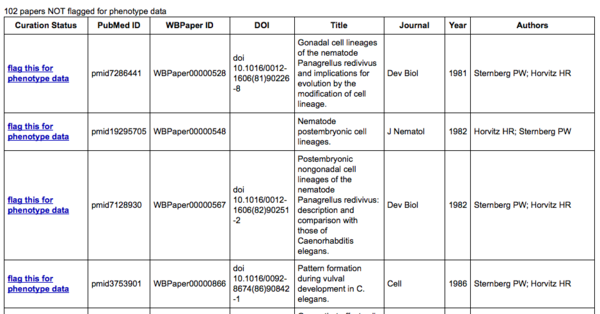
The publication summary page consists of two tables: the top table lists papers for which WormBase data-flagging pipelines have predicted the presence of phenotype data:
There are two distinct pipelines that WormBase uses to detect and curate phenotypes: one for alleles and transgenes and another for RNAi phenotypes. Therefore, there are two indicators as to the status of each paper with regards to phenotype curation. The top status indicator reports the status with regards to allele and transgene phenotype curation and may read "flag this for Allele-Transgene phenotype data", "Allele-Transgene Curation in progress", "Allele-Transgene WormBase curated", or "Allele-Transgene Needs curation". Similarly, the bottom status indicator reports the status of the paper with regards to RNAi phenotype curation and may read "flag this for RNAi phenotype data", "RNAi Curation in progress", "RNAi WormBase curated", or "RNAi Needs curation". The "flag this for ... phenotype data" status indicates that our pipelines have not predicted this paper to contain that type of phenotype data; clicking on the link will flag the paper for phenotype data so that a WormBase curator can add it to the list of papers to curate. The "... Curation in progress" indicates that one or more community curators have submitted phenotype data for this paper but that the curation has not yet been completed by a WormBase curator. The "... WormBase curated" indicates that the paper has been completely curated (or is currently being curated) by a WormBase curator. The "... Needs curation" indicates that the paper has been positively identified to be reporting phenotype data, but no curation has begun. Clicking on the "Needs curation" link will direct you to back to the Phenotype form with your name, e-mail address (if already provided), and PubMed ID of that paper pre-populated into the appopriate fields of the form:
The bottom table lists papers that have not been flagged for phenotype data.
In the bottom table of the publication summary page, clicking on "flag this for phenotype data" will notify a WormBase curator that this paper has phenotype data and should be added to the list of papers considered as having phenotype data. Once clicked, you will be directed to a flagging confirmation page:
For papers that do not have a PubMed ID, a digital object identifier (D.O.I.) may be entered instead. In this case, the term information box will indicate that the ID is not recognized:
but please continue with your submission, and a WormBase curator will contact you if there are any questions.
Genetic Perturbation(s)
Each submission of data via this phenotype form requires at least one genetic perturbation to be indicated as responsible for the phenotype (observed or not). There are three types of genetic perturbation info that the form will accept: RNAi target genes, alleles, and transgenes.
RNAi target gene
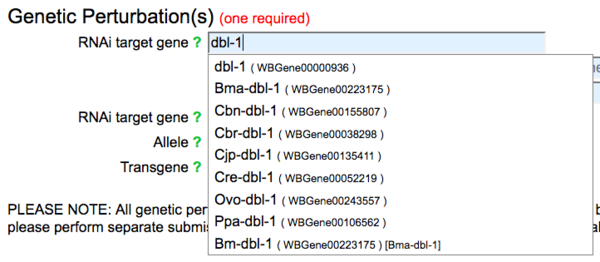
An RNAi target gene is required for submission of RNAi-phenotype data. The "RNAi target gene" field accepts recognized gene names using an autocomplete feature, suggesting genes as you type:
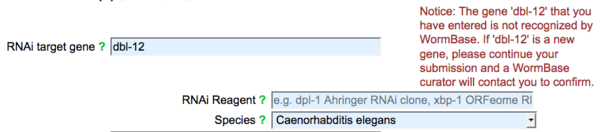
You may select a gene from the suggested list, or if your gene is not listed (not recognized by WormBase) enter the name of your gene and a notice will appear indicating that the gene name is not recognized, but to proceed with your submission and that a WormBase curator will contact you to confirm the new gene name:
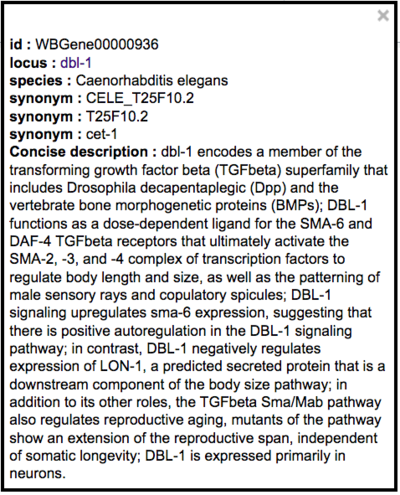
If a recognized gene name is selected from the autocomplete suggestions, a term information box will appear in the upper right corner of the screen displaying basic information about the gene with a link (locus name is hyperlinked) to the respective WormBase gene page:
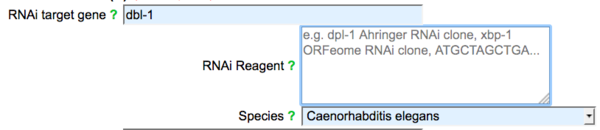
Once a gene has been selected (or typed in), an "RNAi reagent" field and a "Species" field will appear:
You may enter a library name and gene name (e.g. "Ahringer dpl-1 clone"), the commercial clone well location (e.g. "Source BioScience II-6H13") or the sequence of the RNAi clone (e.g. "ACGTATGCTAG...."). Because RNA interference experiments can now be carried out in several species, the form has a species selection to specify which species the RNAi experiment(s) was/were carried out in, with a default to Caenorhabditis elegans. Note that as soon as an entry is made in the "RNAi target gene" field, an additional "RNAi target gene" field will appear, allowing submission of multiple RNAi target genes simultaneously. If an entry is made in the second RNAi target gene field (or more than one genetic perturbation is entered in general), a notice will appear asking the submitter to indicate if the multiple genetic perturbations represent part of a complex genotype (single phenotype experiment) or multiple independent phenotype experiments (each with the indicated phenotype); see below.
Allele
The allele name is required for submission of allele-phenotype data. You may enter a WormBase-recognized allele name or the name of an allele that WormBase has not yet cataloged. Start typing an allele name and suggestions will appear below the field:
Once an allele is selected from the drop down list of WormBase-recognized alleles, a term information box will appear in the upper right-hand corner of the screen with additional information about this allele:
This term information box displays links to the WormBase variation/allele page (via the "Click here..." link as well as the WBVar ID) and the WormBase gene page for the affiliated gene.
A notice will appear to the right of the field to indicate if an allele name that has been entered is not recognized by WormBase:
If you are entering a "new" allele name, click anywhere outside the field to set the name, and continue with your submission. An allele curator will be notified and may contact you to confirm the existence of this allele. Note that as soon as an entry is made in the "Allele" field, an additional "Allele" field will appear, allowing submission of multiple alleles simultaneously. If an entry is made in the second Allele field (or more than one genetic perturbation is entered in general), a notice will appear asking the submitter to indicate if the multiple genetic perturbations represent part of a complex genotype (single phenotype experiment) or multiple independent phenotype experiments (each with the indicated phenotype); see below.
Transgene
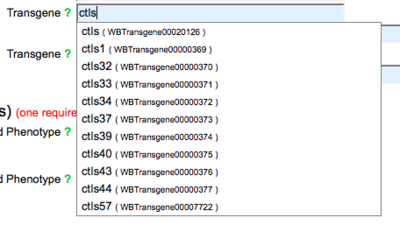
A transgene name is required for submission of transgene-phenotype data. You may enter a WormBase-recognized transgene name or the name of an transgene that WormBase has not yet cataloged. Start typing a transgene name and suggestions will appear below the field:
Once a transgene is selected from the drop down list of WormBase-recognized transgenes, a term information box will appear in the upper right-hand corner of the screen with additional information about this transgene:
This term information box displays links to the WormBase transgene page (via the WBTransgene ID).
Once a transgene has been selected/typed, a "Gene Causing Phenotype" field will appear to enter the gene in the transgene that is most likely responsible for the phenotype observed (e.g. overexpressed gene):
The "Gene Causing Phenotype" field has an autocomplete feature similar to the "RNAi target gene" field, allowing recognized gene names as well as unrecognized (new) gene names with a notice similar to that of the "RNAi target gene" field. A notice will appear to the right of the field to indicate if a transgene name that has been entered is not recognized by WormBase:
If you are entering a "new" transgene name, click anywhere outside the field to set the name, and continue with your submission. A curator will be notified and may contact you to confirm the existence of this transgene. Note that as soon as an entry is made in the "Transgene" field, an additional "Transgene" field will appear, allowing submission of multiple transgenes simultaneously. If an entry is made in the second Transgene field (or more than one genetic perturbation is entered in general), a notice will appear asking the submitter to indicate if the multiple genetic perturbations represent part of a complex genotype (single phenotype experiment) or multiple independent phenotype experiments (each with the indicated phenotype); see below.
Entering multiple genetic perturbations
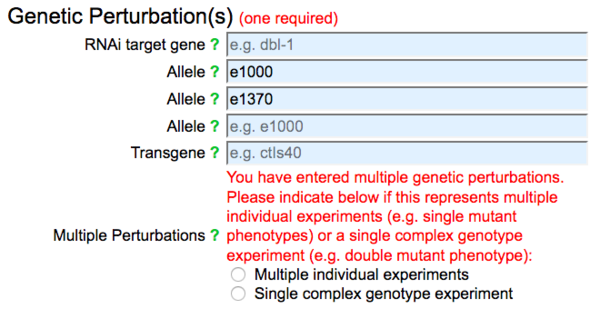
For convenience, the phenotype form allows data submitters to submit multiple genetic perturbations simultaneously in one phenotype data submission. This way, contributors can indicate, in one submission, that multiple experiments each with distinct alleles or RNAi knockdowns, for example, each resulted in the observed phenotype(s) compared to control. Alternatively, the intention may be to indicate that multiple genetic perturbations were simultaneously introduced into some genetic background and a phenotype was observed for a double mutant, for example. To distinguish between these two interpretations, as soon as two or more genetic perturbations are entered into the form, a notice appears asking the user to specify which of these two situations applies for the multiple genetic perturbations entered:
If each genetic perturbation entered represents an individual, independent phenotype experiment, each resulting in the observed phenotype(s), select (click on) the radio button to the left of "Multiple individual experiments", so that the annotation can be properly entered into the WormBase database. On the other hand, if the multiple genetic perturbations entered represents a single complex genotype experiment (e.g. a double or triple mutant phenotype) in which all perturbations were simultaneously assessed for a phenotype in a single organism or isogenic population, select (click on) the radio button to the left of "Single complex genotype experiment".
Note that entering multiple perturbations in a single submission will prevent the form from accepting entries in the "Inheritance Pattern" or "Mutation Effect" fields in the Optional section (see below).
Phenotype(s)
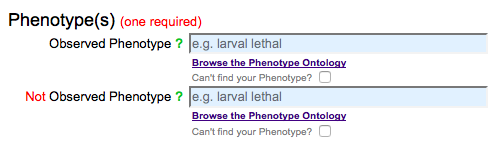
By default, phenotype fields are displayed in the form for both observed and NOT observed phenotypes:
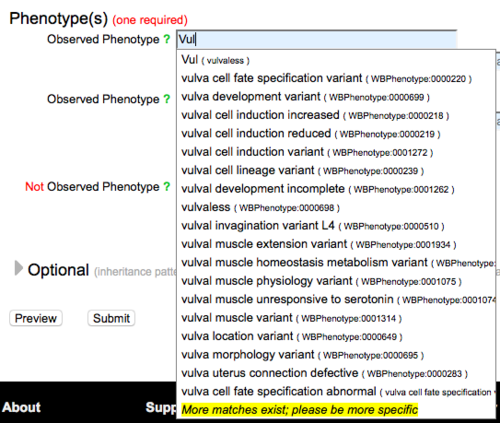
The "Not Observed Phenotype" field is reserved for phenotypes that were assayed for but were not observed (were wild type) for the RNAi clone(s), allele(s), or transgene(s) entered into the form. To enter a phenotype in either field, start typing a phenotype term or a term related to your phenotype and an auto-complete drop down list of official WormBase Phenotype Ontology (WPO) terms that match your entry will appear below the field:
As indicated in the example image above, you may enter a three-letter code for a phenotype which will be matched to phenotype term synonyms, e.g. Let, Lvl, Vul, Muv, Pvl, Gro
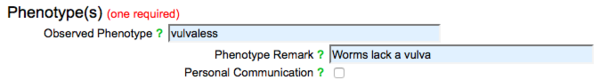
Select a term from the list of WPO terms. Once a phenotype has been selected, enter any pertinent comments about the phenotype with respect to the RNAi clone(s), allele(s), and transgene(s) entered above into the "Phenotype Remarks" field:
Note the presence of a "Personal Communication" checkbox which, if checked, indicates to WormBase that this phenotype was observed independently of any publication and should be considered a personal communication from the submitter (i.e. the paper ID will be ignored for this phenotype).
For WPO terms selected in the phenotype fields, a term information panel will appear in the upper right corner of the page displaying additional information about the phenotype selected as well as links to the WormBase phenotype page showing RNAi clones, alleles, and transgenes known to exhibit this phenotype:
If you cannot find your phenotype in the list of suggested terms, you may browse the WPO using the WormBase Ontology Browser (WOBr). To use WOBr and view the ontology, click on "Browse the Phenotype Ontology" below either of the phenotype fields to be directed to the WormBase Ontology Browser page:
Click on the "+" symbol to the left of the root term "Variant" and subsequent terms to open up the Phenotype Ontology and begin browsing:
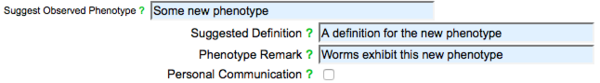
If you still cannot find an appropriate phenotype term, click on the check-box to the right of the text "Can't find your Phenotype?" to open up the "Suggest Phenotype" fields. Enter the name and definition of a suggested new phenotype term in the indicated fields:
A WormBase curator will be notified and will confirm the new phenotype.
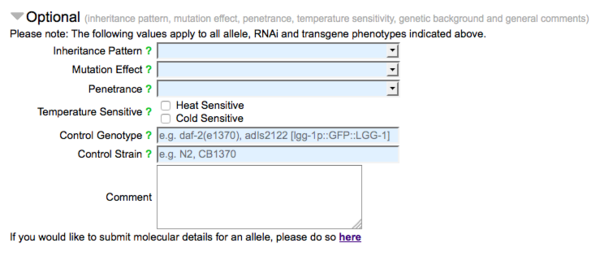
Optional Fields
Below the mandatory fields of the form are a series of fields for optional data, mostly pertaining to the nature of the allele with respect to the phenotype(s) indicated:
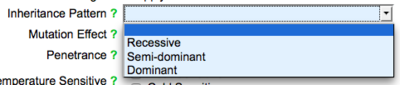
The first field is the "Inheritance Pattern" field, where users may use a drop down list to indicate whether the allele exhibits a recessive, dominant, or semi-dominant inheritance pattern:
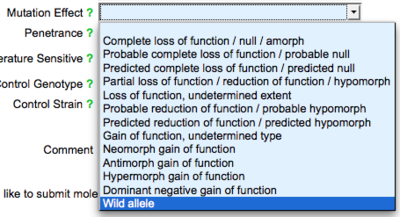
The next field is the "Mutation Effect" field, where users may use a drop down list to indicate the effect of the mutation on the phenotype, e.g. loss of function, gain of function, null, etc. :
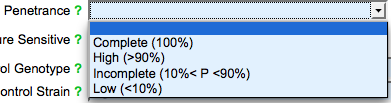
The last drop down list field in the Optional section is the "Penetrance" field, where users may indicate the degree of penetrance observed for the phenotype indicated in the top section of the form:
Below the "Penetrance" field are two check boxes where users may indicate whether the allele is temperature sensitive (heat or cold sensitive) with respect to the phenotype. Submitters are encouraged to specify the genetic background of the experiment either by typing the genotype into the "Control Genotype" field or by entering the name of the strain background in the "Control Strain" field. Lastly, there is a general comments field where submitters may indicate to WormBase curators any additional information important for the annotation:
Preview and Submission
Preview
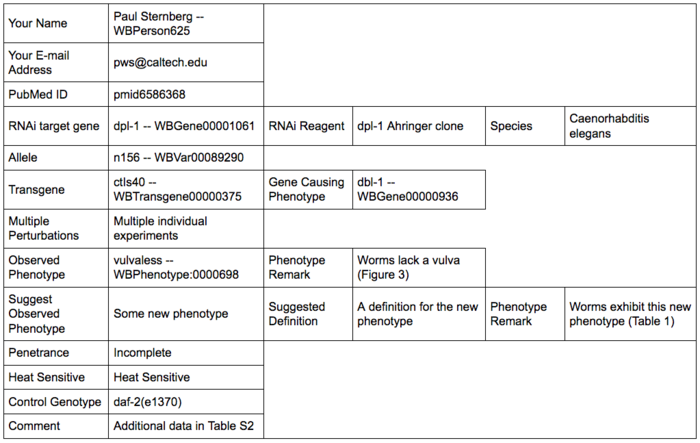
Once all desired fields have been filled, click on the "Preview" button to see a preview of the data submission and check that the data was entered and recognized properly:
Submission
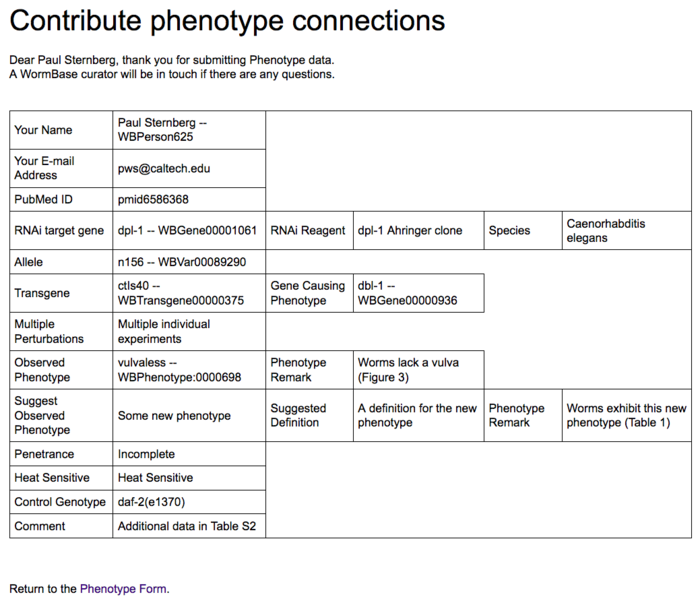
When ready, click the "Submit" button and the data will be submitted to the Postgres database. The user will be directed to a submission confirmation page:
and will receive an e-mail summarizing the content of the submission:
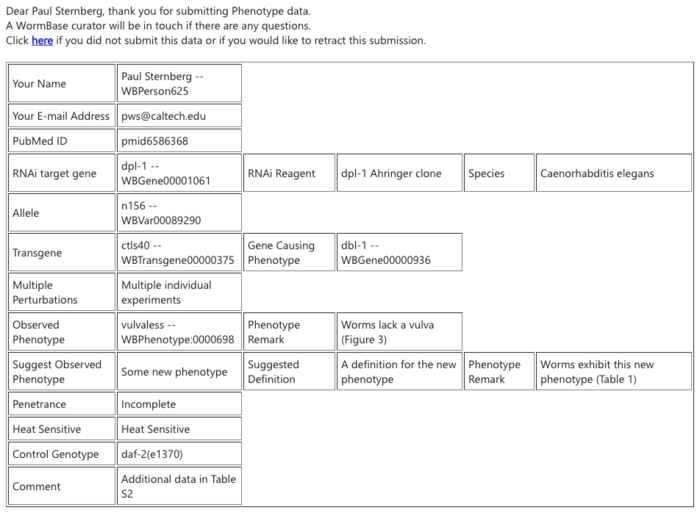
If the submission was unauthorized or accidental, recipients may click on the "Click here if you did not submit this data or would like to retract this submission" to flag the submission for omission. Clicking on this link will direct to a confirmation page:
WormBase Internal Documentation
Form population of Postgres and OA
Each allele-phenotype pair and transgene-phenotype pair submitted via the Phenotype form will be submitted to the Phenotype OA and Postgres as a single entry with a unique Postgres ID (PGID). This means that multiple phenotypes (observed or not) submitted for a single allele or transgene via the form will be entered as separate PGIDs but all with the same shared fields, including community curator name and ID, paper ID and all optional fields. The fields of the form correspond to the Phenotype OA fields in the following manner:
Phenotype Form Field --> Phenotype OA Field (Postgres table name)
Phenotype OA - ALL TABS
- Observed Phenotype --> Phenotype (app_term) (NOT toggled OFF) (app_not)
- Not Observed Phenotype --> Phenotype (app_term) (NOT toggled ON) (app_not)
- Phenotype Remark --> Phenotype Remark (app_phen_remark)
Phenotype OA TAB 1
- N/A --> Curator (app_curator) defaults to "Community Curator (WBPerson29819)" for every submission
- PubMed ID --> Pub (as WBPaper ID) (app_paper)
- Allele --> Variation (app_variation)
- Transgene --> Transgene (app_transgene)
- Control Strain --> Strain (app_strain)
- Comment (Optional section) --> Object Remark (app_obj_remark) [not dumped]
Phenotype OA TAB 2
- Suggested Observed Phenotype --> Suggested (app_suggested) (NOT toggled OFF) (app_not)
- Suggested Not Observed Phenotype --> Suggested (app_suggested) (NOT toggled ON) (app_not)
- Suggested Definition --> Suggested Definition (app_suggested_definition)
Phenotype OA TAB 4
- Inheritance Pattern --> Allele Nature (app_nature)
- Mutation Effect --> Functional Change (app_func)
- Penetrance --> Penetrance (app_penetrance)
- Heat Sensitive --> Heat Sensitive (app_heat_sens)
- Cold Sensitive --> Cold Sensitive (app_cold_sens)
Phenotype OA TAB 5
- Control Genotype --> Genotype (app_genotype)
Phenotype OA TAB 6
- Your Name --> Community Curator (app_communitycurator)
- Your Name --> Person (app_person) for each personal communication
- Your E-mail Address --> Community Curator Email (app_communitycuratoremail)
- PubMed ID --> Unregistered Paper (if unrecognized) (app_unregpaper)
- Allele --> Unregistered Variation (if unrecognized or entire genotype if multiple perturbations entered for a single complex genotype experiment) (app_unregvariation)
- Transgene --> Unregistered Transgene (if unrecognized) (app_unregtransgene)
- "NO DUMP" (app_nodump) and "Needs Review" (app_needsreview) toggled ON by default for every submission
For RNAi-phenotype pairs, data will be submitted to the RNAi OA as follows:
Phenotype Form Field --> RNAi OA Field (Postgres table name)
RNAi OA - ALL TABS
- Observed Phenotype --> Phenotype Observed (rna_phenotype) (NOT toggled OFF) (rna_phenotypenot)
- Not Observed Phenotype --> Phenotype Observed (rna_phenotype) (NOT toggled ON) (rna_phenotypenot)
- Phenotype Remark --> Phenotype Remark (rna_phenremark)
RNAi OA TAB 1
- N/A --> Curator (rna_curator) defaults to "Community Curator (WBPerson29819)" for every submission
- PubMed ID --> Paper (as WBPaper ID) (rna_paper)
- RNAi target gene --> Remark (rna_remark) as "<WBGene ID>"
- RNAi reagent --> DNA text (rna_dnatext)
- RNAi species --> Species (rna_species)
- Control Strain (Optional section) --> Strain (rna_strain)
- Control Genotype (Optional section) --> Genotype (rna_genotype)
- Comment (Optional section) --> Remark (rna_remark)
RNAi OA TAB 2
- Suggested Observed Phenotype --> Phenotype Suggestion (rna_suggested) (NOT toggled OFF) (rna_phenotypenot)
- Suggested Not Observed Phenotype --> Phenotype Suggestion (rna_suggested) (NOT toggled ON) (rna_phenotypenot)
- Suggested Definition --> Suggested Definition (rna_suggested_definition)
- Penetrance (Optional section) --> Penetrance (rna_penetrance)
RNAi OA TAB 4
- Your Name --> Community Curator (rna_communitycurator)
- Your Name --> Person Evidence (rna_person) for each personal communication
- Your E-mail Address --> Community Curator Email (rna_communitycuratoremail)
- PubMed ID --> Unregistered Paper (if unrecognized) (rna_unregpaper)
- "NO DUMP" (rna_nodump) and "Needs Review" (rna_needsreview) toggled ON by default for every submission
Currently (as of February 2018), all submissions through the Phenotype form will send an e-mail to the submitter and to Chris Grove and Gary Schindelman and the "NO DUMP" and "Needs Review" toggles will be turned ON by default in the respective OA's. Community curation submissions through this form can be queried by querying for the "Community Curator (WBPerson29819)" and those in need of review can be queried via the "Needs Review" toggle. A history of all submissions to the form can be viewed at:
http://tazendra.caltech.edu/~azurebrd/cgi-bin/data/phenotype_history.html
Code documentation
Files on Mangolassi/Tazendra required for the Phenotype form:
- CGI script (/home/azurebrd/cgi-bin/forms/phenotype.cgi)
- Javascript (/home/azurebrd/public_html/javascript/phenotype.js)
- Images file (/home/azurebrd/public_html/images)
- History file (/home/azurebrd/public_html/cgi-bin/data/phenotype_history.html)