Contributing Allele Phenotype Connections
The following documentation explains the use and backend for the Allele-Phenotype form found here:
http://tazendra.caltech.edu/~azurebrd/cgi-bin/forms/allele_phenotype.cgi
Watch a short video tutorial on YouTube!!!
https://www.youtube.com/watch?v=_gd87S1h3zg&feature=youtu.be
Contents
User Guide
Finding the Allele-Phenotype form
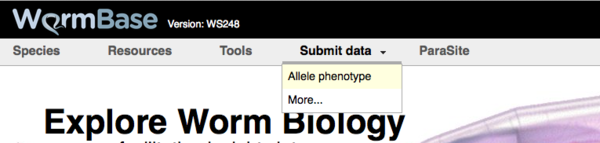
There are two main ways to reach the Allele-Phenotype form from the WormBase homepage. First, and most directly, you can click on the "Allele-Phenotype" option under the "Submit data" menu header at the top of the WormBase homepage:
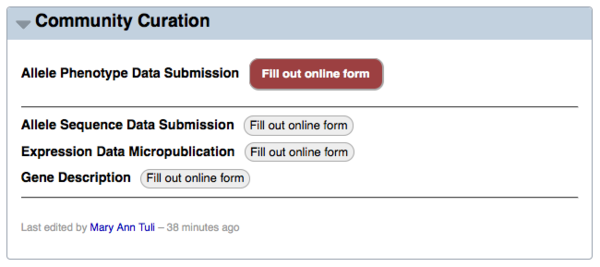
Second, the data submission summary page (accessible by clicking on the "Submit data" menu header at the top of the WormBase homepage) has a link to the form via the red "Fill out online form" button to the right of "Allele Phenotype Data Submission":
Overview
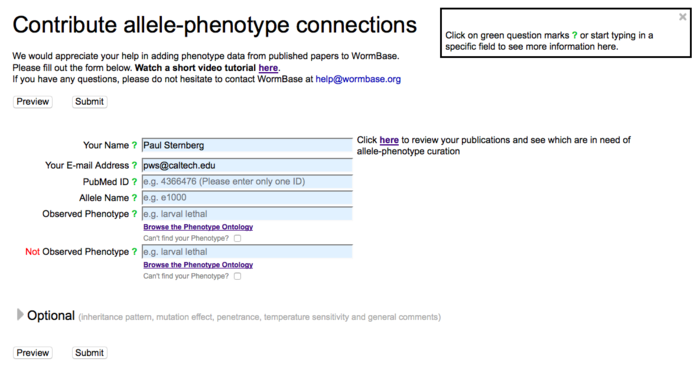
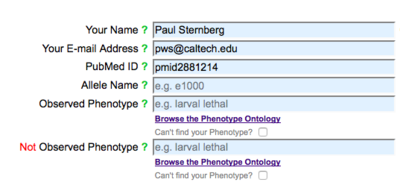
The first part of the Allele-Phenotype form carries all of the fields that are required for data submission: a submitter's name and e-mail address, a publication's PubMed ID, an allele name, and a phenotype (observed or not observed):
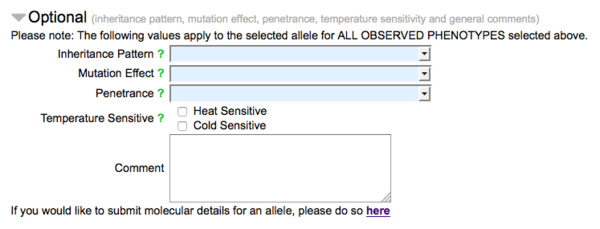
The second part of the form (made visible by clicking on the header "Optional") carries fields for optional data, pertaining to the nature of the allele with respect to the observed phenotypes indicated in the form:
A drop down list of options for inheritance pattern (e.g. dominant, recessive), mutation effect (e.g. loss of function, null, gain of function) and penetrance (roughly categorized as low, incomplete, high, and complete) are available. Check boxes are also available to indicate the temperature sensitivity of the allele. There is also a field for general comments for the submission. The bottom of the "Optional" section has a link to the WormBase Allele-Sequence form, by which sequence details of alleles may be reported.
Required information
The form minimally requires a submitter's name and e-mail address, a publication's PubMed ID, an allele name, and a phenotype (observed or NOT observed).
Your Name & E-mail Address
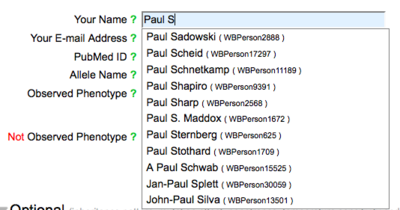
The name field has an autocomplete functionality, requiring the selection of a name from the list of registered WormBase Persons.
If your name is not registered with WormBase, click on the green question mark to the left of the name field:
to open up the help text in the term information box in the upper right-hand corner of the screen and click on the "Person Update Form" link to register.
Once a name has been selected, a term information box will appear in the upper right-hand corner of the page with additional information about this WormBase Person:
including a link to their WormBase Person page, as well as a link to the WBPerson publication summary page (see below). If WormBase has your e-mail address on file, it will automatically be filled into the e-mail field, otherwise you may add (or correct) your e-mail address manually.
PubMed ID
This form requires the entry of a PubMed ID (or D.O.I. if no PubMed ID exists) for the publication in which the allele-phenotype data was originally reported. PubMed IDs may be entered manually with or without the "pmid" prefix, e.g. "pmid6616618" or simply "6616618". Once an ID has been entered and recognized, a term information box will appear in the upper right-hand corner of the screen with additional information about the paper including title, journal, year, and authors list:
To assist in the look-up of PubMed IDs relevant to you, once your name has been selected a notice will appear to the right of the name field with a link to a summary of your publications and their associated PubMed IDs:
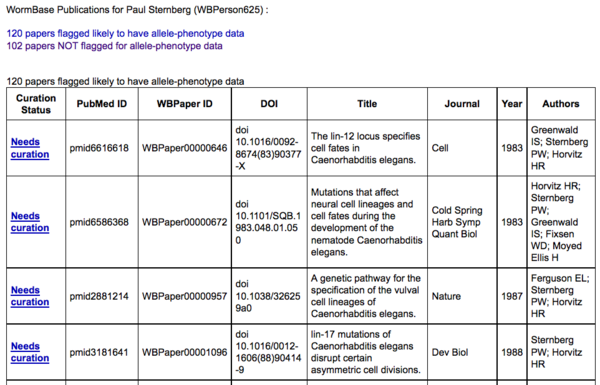
The publication summary page consists of two tables: the top table lists papers for which WormBase data-flagging pipelines have predicted the presence of allele-phenotype data:
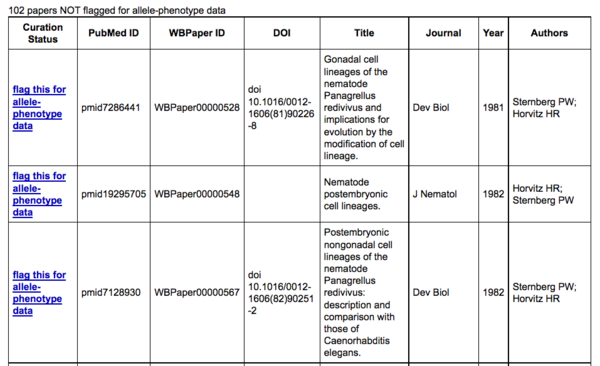
and the bottom table lists papers that have not been flagged for allele-phenotype data.
The top table lists the curation status of papers for which WormBase data-flagging pipelines have predicted the presence of allele-phenotype data, indicating that the paper either "Needs curation" (i.e. there are no allele-phenotype annotations for this paper), has "Curation in progress" (i.e. there is at least one annotation from a community curator using this form) or is "WormBase curated" (i.e. this paper has been curated by WormBase curators). Clicking on the "Needs curation" link will direct you to back to the Allele-Phenotype form with your name, e-mail address (if already provided), and PubMed ID of that paper pre-populated into the appopriate fields of the form:
In the bottom table of the publication summary page, clicking on "flag this for allele-phenotype data" will notify a WormBase curator that this paper has allele-phenotype data and should be added to the list of papers considered as having allele-phenotype data. Once clicked, you will be directed to a flagging confirmation page:
For papers that do not have a PubMed ID, a digital object identifier (D.O.I.) may be entered instead. In this case, the term information box will indicate that the ID is not recognized:
but please continue with your submission, and a WormBase curator will contact you if there are any questions.
Allele Name
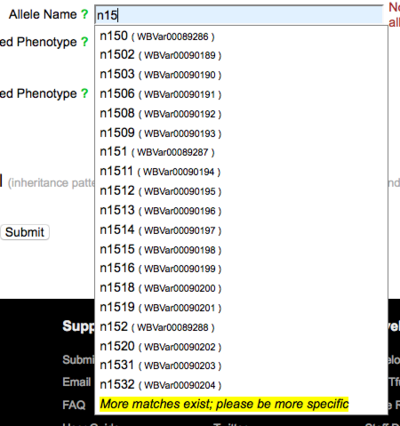
The allele name is required for submission of allele-phenotype data. You may enter a WormBase-recognized allele name or the name of an allele that WormBase has not yet cataloged. Start typing an allele name and suggestions will appear below the field:
Once an allele is selected from the drop down list of WormBase-recognized alleles, a term information box will appear in the upper right-hand corner of the screen with additional information about this allele:
This term information box displays links to the WormBase variation/allele page (via the "Click here..." link as well as the WBVar ID) and the WormBase gene page for the affiliated gene.
A notice will appear to the right of the field to indicate if an allele name that has been entered is not recognized by WormBase:
If you are entering a "new" allele name, click anywhere outside the field to set the name, and continue with your submission. An allele curator will be notified and will contact you to confirm the existence of this allele.
Phenotype
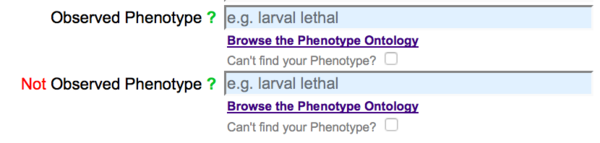
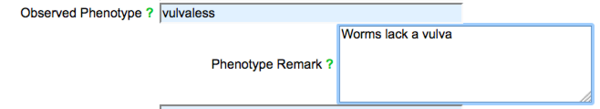
By default, phenotype fields are displayed in the form for both observed and NOT observed phenotypes:
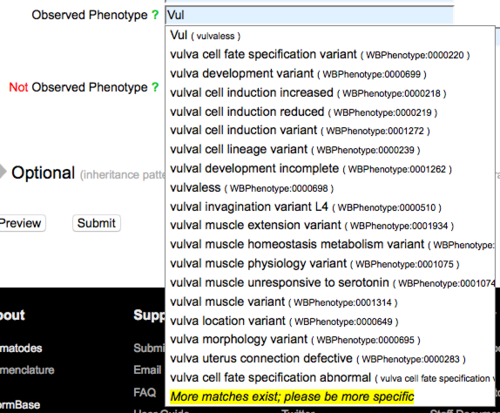
The "Not Observed Phenotype" field is reserved for phenotypes that were assayed for but were not observed (were wild type) for the allele entered into the form. To enter a phenotype in either field, start typing a phenotype term or a term related to your phenotype and an auto-complete drop down list of official WormBase Phenotype Ontology (WPO) terms that match your entry will appear below the field:
As indicated in the example image above, you may enter a three-letter code for a phenotype which will be matched to phenotype term synonyms, e.g. Let, Lvl, Vul, Muv, Pvl, Gro
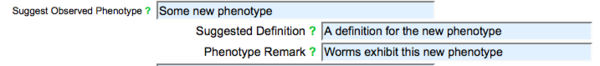
Select a term from the list of WPO terms. If you cannot find your phenotype in the list of suggested terms, you may browse the WPO using the [www.wormbase.org/tools/ontology_browser WormBase Ontology Browser]