Calls
Contents
October 21 2016
1) Replica experiments: -the trehalose paper was sent back because it was repeating published results -> how do we deal with it?. We decided to allow submission of replica experiment as long as is stated. At the moment of submission authors will have to tag the submission:
Section headings New findings Methods & reagents Replication – successful Replication – unsuccessful Negative results
We let the author propose the tag but leave it open to the reviewer to disagree.
Shall we put a limit for submission of replica experiments? -> Yes. How many max? 5? We don’t want to have people exploiting the system and send published results over and over
2) Disclaimer: Make sure that we have a disclaimer, e.g.: 'by publishing i agree to make available the reagents to the community'
3) Add availability of reagents on the file, e.g. N2: CGC
4) Shall we post reviewers comments online? -> topic on the table for future discussion
5) Daniela will contact also Highwire and e-bench press to see what it would entail to work with them
October 12 2016
Agenda
1) Reviewers suggestions and policy (1 vs 2 reviewers) for Nemametrix submissions.
- We want to upload 4 nemametrix micropubs by the October 26th deadline
- Need to send out to reviewers ASAP
- To whom and how many?
2) Collaboration with the Coko foundation. [1]
- They are happy to support us and are developing tools that perfectly fit with our project
- They can write LoS or write grant together with us (Software is under MIT license) -> what do we want to do
3) Format of the micropub journal pages [2]
- does the proposed format look ok?
- this is the page we will send out for review and that will be converted in PDF if the micro is accepted
Minutes
1) Reviewers:
- we will have 1 reviewer and we will leave the option to be anonymous
- we will send the first 2 micropubs to people that are aware of the project
- will also ask authors if they have reviewer's suggestions
- General comment from Tim: we need to put up an advertising show at the IWM
2) Coko
- ok to explore partnering with them
- K and D will set up the whiteboard session in SF to learn more and see what the collaboration will be
- split tasks: we do curation, they do the software -> co-grant?
3) Format of the micro pub
- ok as is now, will try to have a flexible format (one that allows various amounts of narrative)
September 27 2016
Agenda
1) Type of submissions
- For gene expression micropublications, the submission includes:
- reagents used to perform the experiment (transgene, construct, antibody, etc..)
- the read out of the experiment, i.e. image with pattern description and curated metadata
- for phenotype micropublications, Nemametrix has 2 kind of tech notes:
- http://nemametrix.com/trehalose-extends-healthspan-c-elegans/
- http://nemametrix.com/microfluidic-epg-recordings-diverse-nematode-species/
the first one falls into the micropublication scope, the second is more like a Worm Breeder's Gazette article. Discuss.
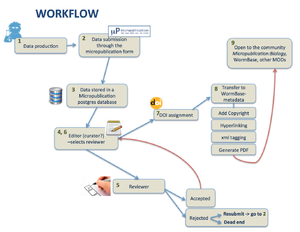
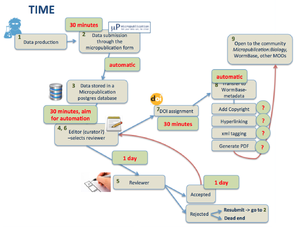
2) Workflow
- from Submission through the form till acceptance by a reviewer we need to track the submission and store metadata in a proper way. Are we going to:
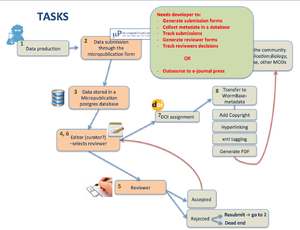
- outsource to e-journal press or similar?
- develop in house? Need developer time, Who is going to do it
3) Editorial Board
- if we start a journal we need an EIC (Paul?) and an editorial board.Who will select and contact the potential editorial board, when.
4) Need alternative title than 'Micropublication: biology' as colons can give problems for downstream XREF (from G3 experience)
Minutes
1) Micro vs milli
- Tim suggests to have narrative-containing Micropubs more similar to a short journal article
- Those could be millipublications as opposed to just data-driven micropublications
- Karen and Daniela will work on different types of Tech notes from Nemametrix and on Worm Breeder's Gazette articles to model that
- multiple micropubs can be put together to have the narrative-containing (hypothesis driven) millipub
- micropubs in the future could be a plug in we give to journal to collect metadata.
2) Workflow. Outsourcing vs in-house
- Todd: outsourcing gives us more expertise but we loose control
- Would be good to have more than one developer involved. Not much resources at the moment
- We will touch base with the Collaborative Knowledge Foundation (CoKo, http://coko.foundation) and see what a collaboration would entail.
- We will continue to develop forms in-house to move on with the project
- we will generate a database with Juancarlos that will store all data but is separate from Postgres used for WB data
3) Editorial board
- PWS can be EIC, need to define scope and build editorial board
4) Title
- We can call it 'Micropublication biology'
- Daniela will check with XREF if calling it 'Micropublication-biology' with a dash -as per Tim's suggestion- will also be an possibility
5) Random
- Tim suggesting to capture crispr reagents -> having them published as micros
6) We will have another call in 2 weeks -October 11 approx- unless we have more news from Nemametrix or there is something compelling to discuss.