Calls
September 27 2016
Agenda
1) Type of submissions
- For gene expression micropublications, the submission includes:
- reagents used to perform the experiment (transgene, construct, antibody, etc..)
- the read out of the experiment, i.e. image with pattern description and curated metadata
- for phenotype micropublications, Nemametrix has 2 kind of tech notes:
- http://nemametrix.com/trehalose-extends-healthspan-c-elegans/
- http://nemametrix.com/microfluidic-epg-recordings-diverse-nematode-species/
the first one falls into the micropublication scope, the second is more like a Worm Breeder's Gazette article. Discuss.
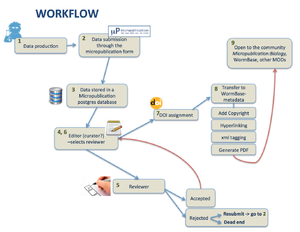
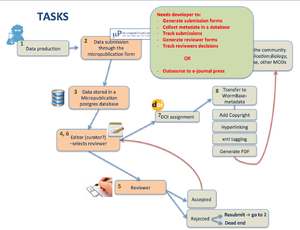
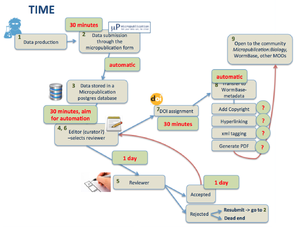
2) Workflow
- from Submission through the form till acceptance by a reviewer we need to track the submission and store metadata in a proper way. Are we going to:
- outsource to e-journal press or similar?
- develop in house? Need developer time, Who is going to do it
3) Editorial Board
- if we start a journal we need an EIC (Paul?) and an editorial board.Who will select and contact the potential editorial board, when.
4) Need alternative title than 'Micropublication: biology' as colons can give problems for downstream XREF (from G3 experience)
Minutes
1) Micro vs milli
- Tim suggests to have narrative-containing Micropubs more similar to a short journal article
- Those could be millipublications as opposed to just data-driven micropublications
- Karen and Daniela will work on different types of Tech notes from Nemametrix and on Worm Breeder's Gazette articles to model that
- multiple micropubs can be put together to have the narrative-containing (hypothesis driven) millipub
- micropubs in the future could be a plug in we give to journal to collect metadata.
2) Workflow. Outsourcing vs in-house
- Todd: outsourcing gives us more expertise but we loose control
- Would be good to have more than one developer involved. Not much resources at the moment
- We will touch base with the Collaborative Knowledge Foundation (CoKo, http://coko.foundation) and see what a collaboration would entail.
- We will continue to develop forms in-house to move on with the project
- we will generate a database with Juancarlos that will store all data but is separate from Postgres used for WB data
3) Editorial board
- PWS can be EIC, need to define scope and build editorial board
4) Title
- We can call it 'Micropublication biology'
- Daniela will check with XREF if calling it 'Micropublication-biology' with a dash -as per Tim's suggestion- will also be an possibility
5) Random
- Tim suggesting to capture crispr reagents -> having them published as micros
6) We will have another call in 2 weeks -October 11 approx- unless we have more news from Nemametrix or there is something compelling to discuss.
October 12 2016
Agenda
1) Reviewers suggestions and policy (1 vs 2 reviewers) for Nemametrix submissions.
- We want to upload 4 nemametrix micropubs by the October 26th deadline
- Need to send out to reviewers ASAP
- To whom and how many?
2) Collaboration with the Coko foundation. [1]
- They are happy to support us and are developing tools that perfectly fit with our project
- They can write LoS or write grant together with us (Software is under MIT license) -> what do we want to do
3) Format of the micropub journal pages [2]
- does the proposed format look ok?
- this is the page we will send out for review and that will be converted in PDF if the micro is accepted
Minutes
1) Reviewers:
- we will have 1 reviewer and we will leave the option to be anonymous
- we will send the first 2 micropubs to people that are aware of the project
- will also ask authors if they have reviewer's suggestions
- General comment from Tim: we need to put up an advertising show at the IWM
2) Coko
- ok to explore partnering with them
- K and D will set up the whiteboard session in SF to learn more and see what the collaboration will be
- split tasks: we do curation, they do the software -> co-grant?
3) Format of the micro pub
- ok as is now, will try to have a flexible format (one that allows various amounts of narrative)