Administration:Installing WormMine
How to set up a development instance of WormMine
Contents
- 1 Quick Overview
- 2 Requirements
- 3 Installation / configuration
- 4 Get production database
- 4.1 Build a new production database
- 4.2 Instantiate database dump
- 4.3 Copy an existing database
- 4.4 Migrating from the unmerged to staging branch
- 4.5 Migrating to production machine
- 4.6 Create userprofile database
- 5 Attach to WormBase instance
- 6 Upgrading between releases
- 7 Development in a Nutshell
Quick Overview
Work towards staging and production should be carried out as "intermine" user (sudo su - intermine).
- InterMine machines:
- production: 206.108.125.166
- development: 206.108.125.174
- Important directories:
- database backup: /nfs/wormbase/wormmine/database_dumps
- in intermine directory:
- datadir: holds data files
- build_config: config files for build
- redeployment: mine update build
Login information is available in a mine's ~/.intermine/wormmine.properties file.
Requirements
Hardware
Linux
- 8 cores
- 24GB RAM
- ~ 1TB storage
Software
Necessary software and versions:
| Software | Minimum Version | Purpose |
|---|---|---|
| Git | 1.7 | check out and update source code |
| Java SDK | 6.0 | build and use InterMine |
| Ant | 1.8 | invokes the InterMine build |
| Tomcat | 6.0.29 | website |
| PostgreSQL | 8.3 | database |
| Perl | 5.8.8 | run build scripts |
Installation / configuration
Environment Setup
Add "intermine" user:
sudo adduser intermine
Password available from Joachim (later: Abigail, Todd).
Dependencies
Git
Install the command line tool:
$ sudo apt-get install git-core
Configure your user and email:
$ git config --global user.name "Name Surname"
$ git config --global user.email "your.email@gmail.com"
Java
Download here.
Since InterMine can be memory intensive, it's helpful to pass environment variables to ant through the ANT_OPTS variable.
TODO: figure out the real reason why these parameters should be used. They have little to do with memory usage, but appear to optimize for throughput.
$ export ANT_OPTS="-server -XX:MaxPermSize=256M -Xmx1700m -XX:+UseParallelGC
-Xms1700m -XX:SoftRefLRUPolicyMSPerMB=1 -XX:MaxHeapFreeRatio=99"
Ant
Refer to ant's manual for installation instructions.
Tomcat
# Assuming you cloned website-intermine first: # git clone https://github.com/WormBase/website-intermine.git ./website-intermine/scripts/install_tomcat.sh 7.0.47
Note: the version number (7.0.47) might be out-of-date. To get a listing of currently available versions, type:
./website-intermine/scripts/install_tomcat.sh
Starting Tomcat
cd apache-tomcat-TOMCATVERSION ./bin/startup.sh
Stopping Tomcat
cd apache-tomcat-TOMCATVERSION ./bin/shutdown.sh
Configuring an Alternative HTTP Port
vim apache-tomcat-TOMCATVERSION/conf/server.xml
Replace the port number in this context:
<Service name="Catalina">
<Connector port="YOURPORT" protocol="HTTP/1.1"
connectionTimeout="20000"
redirectPort="8443"
URIEncoding="UTF-8" />
...
For user-specific port numbers see the Google Doc "Developer Resources at WormBase".
PostgreSQL
Refer to InterMine PostgreSQL installation guide
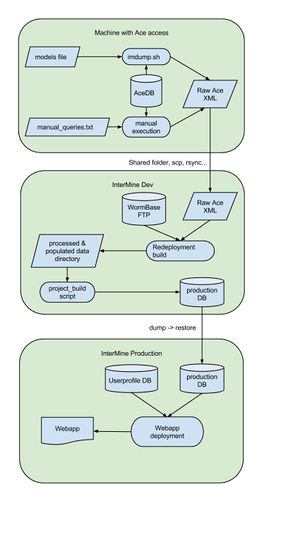
WormMine Workflow
CREATE ROLE intermine_admin WITH SUPERUSER LOGIN PASSWORD 'SECRET';
Perl
Refer to InterMine Perl installation guide
Download and Install WormMine
git clone https://github.com/WormBase/website-intermine.git ./website-intermine/scripts/install_intermine.sh ENVIRONMENT
ENVIRONMENT is either of the following options:
- production
- staging
- development
Create properties file
- Create ~/.intermine directory.
- Copy the sample properties file as
~/.intermine/wormmine.properties - Fill in placeholders as follows:
- <POSTGRES USER PASSWORD>: postgres password for intermine user
- <TOMCAT USER PASSWORD>: tomcat password for intermine user
- <SERVER PUBLIC BASE URL>: Base url of your web server, including port. Sample: http://123.456.789.123:8080
- <CREATE WM ADMIN USERNAME/PASSWORD> create the primary admin account
- <EMAIL ADDRESS TO SEND HELP EMAILS FROM>: your server should be configured to send emails from this address. This will send users password reset emails and the like.
- <HELP REQUESTS ARE SENT HERE>: can be same address as above, this is where input from the InterMine help form gets sent.
Get production database
Build a new production database
Custom WormBase loading processes
Protein Fasta
- custom loader
WormBaseProteinFastaLoaderTask.java
- puts the first ID of the title row into both Protein.primaryIdentifier and Protein.primaryAccession, and the second ID into protein.secondaryIdentifier
GFF3
- custom loader
WormbaseGff3CoreGFF3RecordHandler.java
- when processing transcript record it creates a reference to it's parent gene
- when processing coding sequence record it creates a reference to it's parent transcript
XML
- custom loader
WormbaseAcedbConverter.java
- can load any XML, not ace specific
- loads intermine class fields as mapped by mapping file
Mapping file format:
# this is a comment, comment lines are ignored # regular annotation, this field will be filled in # with the value of the evaluated xpath primaryIdentifier = /Variation/text()[1] # returns true if xpath returns any nodes at all, # useful for data contained in ace tags themselves if.naturalVariant = /XPATH/... # type casting allowed, this example will add the value as a # Phenotype record (Phenotype)parents = /XPATH/...
The build requires:
- Fasta
- GFF3
- GO ontology
- GO association
- Ace XML files
All of these are retrieved from the FTP site, except for the Ace XML files.
Generate data files
On dev.wormbase.org, run:
./website-intermine/scripts/dump_ace.sh DUMPPATH
The instructions below will be obsolete, once the automatic execution of the script above is confirmed to work.
These must be created on a machine with a tace instance, and stored in an accessible location.
Generate Ace XML files
Both the blanket dump and manual queries must be run and saved to XML to represent all covered types
Blanket dump
On machine with Ace instance.
- Download the website-intermine repository as described above
- Navigate to acedb-dev/acedb
website-intermine]$ cd acedb-dev/acedb/
imdump.sh is a shell script which generates XML files for each species in model, into the supplied destination directory. The intermine machine must have access to the directory.
Run it with: any folder can be used as the Ace XML dump location
> ./imdump.sh <ACE XML DUMP>
- It requires the Ace DB database location, is none is supplied through the $ACEDB environment variable, it will search
/usrfortaceand use<it's grandparent directory>/wormbase
/usr/local/wormbase/wormmine/xmldumps/
[19:20|jdmswong@ip-10-35-66-254|acedb]$ ./imdump.sh /usr/local/wormbase/wormmine/xmldumps/
Did not specify AceDB dir in $ACEDB. Searching for tace...
ACEDB set to: /usr/local/wormbase/acedb/wormbase
Species
... done.
Gene
... done.
... and so on
If the directory passed into imdump.sh contains a trailing slash, script will not function correctly.
If XML files for each type in models has been created, then the script has executed successfully. The beginning and ends of each file may be checked manually to be sure.
If the error displayed in the output below is encountered, Ace is probably being disrupted and the command should be re-ran
jbaran@ip-10-35-66-254:~/src/website-intermine/acedb-dev/acedb$ ./imdump.sh /usr/local/wormbase/website/jbaran/acedbdump
Did not specify AceDB dir in $ACEDB. Searching for tace...
ACEDB set to: /usr/local/wormbase/acedb/wormbase
Species
... done.
Gene
... done.
Transcript
... done.
CDS
... done.
Variation
./imdump.sh: line 40: 19068 Killed $ACEDB_BIN/tace "$ACEDB" > /dev/null <<EOF
wb
find ${model}
show -x -f "$dumpdir/$model.xml"
EOF
... done.
Manual queries
Not all records are desired for some types. In these cases specialized ace queries much be run, a step which is not yet automated.
Commands to generate WS239 build, in website-intermine/acedb-dev/acedb directory:
tace "/usr/local/wormbase/acedb/wormbase" acedb> query find Gene Live acedb> show -x -f <ACE XML DUMP>/Gene.xml acedb> query find Protein Corresponding_CDS acedb> show -x -f <ACE XML DUMP>/Protein.xml acedb> query find CDS Method="curated" acedb> show -x -f <ACE XML DUMP>/CDS.xml acedb> query find Transcript (Gene) acedb> show -x -f <ACE XML DUMP>/Transcript.xml acedb> KeySet-Read species.ace acedb> show -x -f <ACE XML DUMP>/Species.xml
The files generated will reflect the ace queries used to generate them. All species are loaded unless otherwise specified.
Each query in website-intermine/acedb-dev/acedb/manual_queries.txt must be run individually in tace, followed by show -x -f <TYPE>.xml where <TYPE> is the ace type being queried.
These queries represent desirable subsets of those represented types.
Acquire and pre-process data files
Copy the Ace XML over from the shared directory they were saved to from the machine which generated them or use:
rsync <HOST IP>:<HOST MACHINE FILE LOCATION> <TARGET MACHINE IP>:<TARGET MACHINE PATH>
Acquires data files from their appropriate data sources, and pre-processes each one accordingly.
On the InterMine machine, in the intermine directory:
- Navigate to the redeployment folder.
intermine]$ cd redeployment/
- The
update.propertiesfile should contain these two entries:
release = WS239
ace-xml-dir = /nfs/wormbase/wormmine/acedb_dumps/${release}
The release is used to generate strings, and must match the format of the FTP site.
ace-xml-dir is where the build looks for the ace xml files generated above. This has to be set to <TARGET MACHINE PATH> from above.
The build downloads and processes fasta, gff3, go, gaf files. In addition to copying and processing the Ace XML files.
Build configuration
| Property | function |
|---|---|
| datadir | The data directory for WormMine |
| release | WormBase release version to use for paths and filenames |
| backup-dirname | directory to backup old data directory too. |
| genomic-fasta-species-file | species to download and/or process genomic fasta for |
| protein-fasta-species-file | species to download and/or process protein fasta for |
| gff3-species-file | same, for gff3s |
| ace-classes file | ace classes to copy and/or process |
ant -p will display all invokable tasks, available for individual execution.
- Run the build
This will backup the old data directory into backup-dirname, delete it, then download and process all file types.
redeployment]$ ant
To only download and process:
redeployment]$ ant run-all
Size requirements:
- WS239: 70G
- WS240: 75G
Troubleshooting
Malformed JSON string: output produced by "ant run-all"
website-intermine/acedb-dev/intermine/redeployment$ ant run-all
Buildfile: /home/yourusername/src/intermine/redeployment/build.xml
get-assembly-ids:
[exec] malformed JSON string, neither array, object, number, string or atom, at character offset 8424 (before "],\n "full_name..."JSON retrieved from ftp://ftp.wormbase.org/pub/wormbase/releases/WS239/species/ASSEMBLIES.WS239.json
[exec] ) at gen_assemblies.pl line 19.
BUILD FAILED
/home/yourusername/src/intermine/redeployment/build.xml:39: exec returned: 255
Total time: 2 seconds
Fix: Contact Kevin Howe so that the JSON file on the FTP server can be updated. JSON formatting errors occur sometimes when the file is manually edited after its generation.
Build the database
- From the intermine directory, navigate to
wormmine
cd wormmine mkdir /YOURPATH/datadir/entrez-organism
- Run build
wormmine]$ ../bio/scripts/project_build -b -v localhost wormmine_wsVERSION_PATCH
- VERSION: WormBase release version (e.g. WS240)
- PATCH: InterMine build patch, which indicates rebuilds that were requested due to data inconsistencies, data loss, etc. (e.g. 3)
This will run all sources configured in intermine/wormmine/project.xml file.
The project.xml file is described in more detail in the official documentation.
Issues can be addressed on the InterMine developer list: dev (AT) intermine.org
Output file: wormmine_wsVERSION_PATCH.final
Log file: pbuild.log
Troubleshooting
If an exceptions occur, then "pbuild.log" should be checked for a more detailed explanation.
Build Steps (Preliminary Info)
Building the database refers to the process in which the project build script compiles each source together into a production database.
It is unclear which parts of the database are build using "build-db", which cannot recover/be resumed when it fails, and everything coming afterwards.
1. Build DB
Invoked by the -b switch to project_build.
Runs: cd dbmodel ; ant clean build-db
This generates the primary mine model file, and creates a fresh database with the desired schema. To generate the model it merged the model additions for all source types used in the main project.xml file. Ace XML, Gff3, and Fasta sources are the most essential.
Restarting Building from Milestones/Checkpoints
If a build error is encountered, it will appear in the output of the build command as "BUILD FAILED". To restart from the latest checkpoint, run the project build script with the -b (build) flag replaced with -l (recover). If the -b flag is not omitted, then "build-db" will be run again, which can take a long time.
../bio/scripts/project_build -v -l localhost wormmine_dump
This will, instead of rebuilding, attempt to restart by reading the last dump database. Dump databases are created by the build script from sources through specifications in the project.xml through the "dump" attribute. An example:
<source name="wb-acedb-Variation" type="wormbase-acedb" dump="true">
properties go here ...
</source>
The builder will create the <DATABASE NAME>:wb-acedb-Variation database once this source is run. If restarting, the builder will find the most recent of these backup databases, clone it, and resume from there.
About the database
The database name is configured in the properties file as: db.production.datasource.databaseName. Each table represents a class in the model, with additional ones representing many-to-many collections, and various metadata. The InterMine development team does not currently advise for developers to modify the backend database due to many layers of inheritance, although questions may be directed to the InterMine developer list at dev (AT) intermine.org.
Instantiate database dump
Backup archive can be found at
/nfs/wormbase/wormmine/database_dumps
To instantiate a previously built WormMine production database.
- Find your favorite release from WORMMINE DB FTP URL (placeholder, no URL exists)
- Create empty DB
> createdb -U intermine -E SQL_ASCII wormmine
- -U: user set to intermine
- -E: character set used
- Unpack and restore DB
> psql -U intermine -d wormmine -f <WORMMINE RELEASE SQL>
- -U: execute as user
- -d: destination DB
- -f: SQL input file
Copy an existing database
Creates a new database with the contents of an already existing database:
createdb -U intermine -W -T EXISTINGDBNAME NEWDBNAME
Note: the existing database needs to be owned by the "intermine" PostgreSQL user. If that is not the case, then it can be set using the following SQL command:
ALTER DATABASE EXISTINGDBNAME OWNER TO intermine
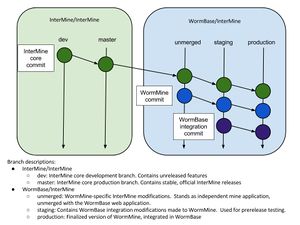
Migrating from the unmerged to staging branch
git merge is used to move the files and changes on the unmerged branch into staging.
[15:32|jdmswong@wb-intermine|webapp]$ git checkout staging Switched to branch 'staging' [15:34|jdmswong@wb-intermine|webapp]$ git merge unmerged --squash ....... [15:36|jdmswong@wb-intermine|webapp]$ git commit
Migrating to production machine
The production database must be moved to the production server to be served by the webapp.
Any paths used in the following commands are arbitrary as long as they are consistent.
Some databases are named along the lines of wormmine-ws239-2. Database names are arbitrary as long as they are referred to correctly in the mine properties file.
Dumping database
On the development machine:
pg_dump -U intermine <DATABASE NAME> -f <PATH TO DUMPFILE>
This created the database dump file.
Transfer this dumpfile to the production machine. Scp is one of the many options: On development machine:
scp <PATH TO DUMPFILE> <REMOTE MACHINE IP>:<REMOTE DUMPFILE PATH>
Restoring database
On production machine:
createdb -U intermine -E SQL_ASCII wormbase-wsVERSION-PATCH pg_restore -U intermine_admin -d wormbase-wsVERSION-PATCH wormbase-wsVERSION-PATCH.final
Note: the psql command below is probably wrong. Will be removed once I can confirm that the pg_restore works.
psql -U intermine -d <DESTINATION DATABASE NAME> -f <REMOTE DUMPFILE PATH>
The database is now instantiated on the production machine
If the following error is encountered, your postgres user must be granted super user privilege.
jbaran@wb-intermine:/nfs/wormbase/archive/jbaran/WS239-2$ psql -U intermine -d joachim-ws239-2 -f wormmine-ws239-2.sql SET SET SET SET SET SET SET psql:wormmine-ws239-2.sql:18: ERROR: must be superuser to create a base type psql:wormmine-ws239-2.sql:27: ERROR: permission denied for language c ALTER FUNCTION psql:wormmine-ws239-2.sql:38: ERROR: permission denied for language c ALTER FUNCTION psql:wormmine-ws239-2.sql:53: ERROR: must be superuser to create a base type ALTER TYPE psql:wormmine-ws239-2.sql:62: ERROR: must be owner of type bioseg psql:wormmine-ws239-2.sql:71: ERROR: permission denied for language c ALTER FUNCTION psql:wormmine-ws239-2.sql:80: ERROR: must be owner of function bioseg_cmp psql:wormmine-ws239-2.sql:89: ERROR: permission denied for language c ALTER FUNCTION psql:wormmine-ws239-2.sql:98: ERROR: must be owner of function bioseg_contained psql:wormmine-ws239-2.sql:107: ERROR: permission denied for language c ALTER FUNCTION psql:wormmine-ws239-2.sql:116: ERROR: must be owner of function bioseg_contains psql:wormmine-ws239-2.sql:125: ERROR: permission denied for language c
Configuring mine to database
On production machine, in
/home/jdmswong/.intermine/wormmine.properties
, set
db.production.datasource.databaseName=<DESTINATION DATABASE NAME>
Create userprofile database
InterMine needs a separate database to track users and their information.
This can be skipped if an existing userprofile database is present.
- Create empty DB
> createdb -U intermine -E SQL_ASCII userprofile-wormmine
- Build the userprofile DB
> cd wormmine/webapp > ant build-db-userprofile
This formats the empty userprofile database for mine use.
About the userprofile database
The database name is set in the properties file as db.userprofile-production.datasource.databaseName. User information is stored in the userprofile table. Tables that begin with "saved" map users to any data they have saved; such as lists, queries, templates, and so on. List data mapping is stores in bagvalues
Launch webapp
- Note: Catalina must be running before you deploy the webapp (about catalina)
- also, you must undeploy any instances that may currently be running
- Navigate to intermine/wormmine/webapp
- Launch webapp:
> ./launch_webapp.sh
- Note: for users who aren't JD, this must be run as
> sudo -ujdmswong ./launch_webapp.sh
This script contains:
ant clean ant -v default remove-webapp release-webapp
Which may be run in sequence instead. These commands clear previous webapp files, remove any existing webapps which may be launched, and compile and release a new webapp.
Test Webapp
You should be able to reach your new instance through <baseurl>/wormmine Webapp is standalone.
About Catalina
Catalina is Tomcat's servlet container. Catalina implements specs for servlet and JSP.
$CATALINA_HOME = /home/jdmswong/website-intermine/software/tomcat/apache-tomcat-6.0.36
Catalina must be running before you deploy the webapp
$CATALINA_HOME/bin/startup.sh
In case of problems with deploying the webapp, try restarting Catalina
$CATALINA_HOME/bin/shutdown.sh $CATALINA_HOME/bin/startup.sh
Managing applications
You can use the Tomcat Web Application manager to view/start/stop/undeploy any applications that may be running.
- Production:
http://206.108.125.166:8080/manager/html - The username and password can be found in
/home/jdmswong/website-intermine/software/tomcat/apache-tomcat-6.0.36/conf/tomcat-users.xml- look for
roles="manager-gui"
- look for
You will need to undeploy /tools/wormmine every time you restart the web app. You can do this by clicking on 'undeploy' from the list of applications.
WebApp Logging
Logs can be found:
intermine/wormmine/webapp/intermine.log <$CATALINA_HOME>/logs
Note:
$CATALINA_HOME = /home/jdmswong/website-intermine/software/tomcat/apache-tomcat-6.0.36
TODO: document which logs record what
Attach to WormBase instance
If you want to enable integration with WormBase, follow these steps:
Checkout merged branch
> git checkout remotes/origin/staging Note: checking out 'remotes/origin/staging'. You are in 'detached HEAD' state. You can look around, make experimental changes and commit them, and you can discard any commits you make in this state without impacting any branches by performing another checkout. If you want to create a new branch to retain commits you create, you may do so (now or later) by using -b with the checkout command again. Example: git checkout -b new_branch_name HEAD is now at 611b791... new tests, shell script to run tests > git checkout -b staging Switched to a new branch 'staging'
Reconfigure properties file
Properties file located ~/.intermine/wormmine.properties (currently /home/jdmswong/.intermine/wormmine.properties/)
- Needs to deploy at tools/wormmine
webapp.path=tools/wormmine
- Update url as appropriate:
- For deployment on staging.wormbase.org:
webapp.baseurl=http://staging.wormbase.org webapp.returnurl=http://staging.wormbase.org/auth/openid?openid_identifier=https://www.google.com/accounts/o8/id&redirect=http://dev.wormbase.org/tools/wormmine/mymine.do# project.sitePrefix=http://staging.wormbase.org/tools/wormmine
- For deployment at www.wormbase.org
webapp.baseurl=http://www.wormbase.org webapp.returnurl=http://www.wormbase.org/auth/openid?openid_identifier=https://www.google.com/accounts/o8/id&redirect=http://www.wormbase.org/tools/wormmine/mymine.do# project.sitePrefix=http://www.wormbase.org/tools/wormmine
Modify wormbase.conf
To enable login system, make sure config flag: wormmine_path = 'tools/wormmine' is uncommented.
Upgrading between releases
This process has not been fully automated yet, and thus requires some manual work.
Upgrade release database
Instructions above
Upgrade wormmine.properties file
The current release uses /home/jdmswong/.intermine/wormmine.properties.
Relevant properties:
# this is the production database the mine will use db.production.datasource.databaseName=wormmine-ws238-3 # This appears at the top next to Version WS project.releaseVersion= 238 IM v1.2.1
Update genomic_model.xml
This is the central model file, used by the webapp process. It must be imported from the machine which produced the production database.
Note: it can be added to the repository, but updates after each test build. This led to extraneous commits and merging conflicts.
In /home/jdmswong/intermine/wormmine/dbmodel/build/model
[17:44|jdmswong@wb-intermine|model]$ rsync 206.108.125.174:/home/jdmswong/idev/wormmine/dbmodel/build/model/genomic_model.xml .
Development in a Nutshell
Please provide a walkthrough on how new AceDB class could be added to WormMine. List which files need to be added (incl. why), which configuration files need to be updated, and how https://github.com/WormBase/intermine/blob/unmerged/build_config/wormbase-acedb/Gene_mapping.properties mappings work.
Property files are projecting columns (?) to Ace values via XPath expressions. It is unclear where the data ends up though.