WormBase-Caltech Weekly Calls December 2017
From WormBaseWiki
Jump to navigationJump to search
December 7, 2017
Next Citace upload
- Submit files to Wen by Jan 16, 2018
- There shouldn't be model changes (that affect Caltech) for this next upload
- Juancarlos goes on vacation Dec 19th
WB Curator candidate
- April, could join WB for 1-2 years
- Could possibly interview next Thursday
- Could curate sequence features, enhancers, alleles/mutations/variations?
- Could start in January
Expression clusters
- Wen and Raymond discussed; will add an "uncertain" tag or something similar
- Tag can be used to filter out datasets that are complex (like mixed cell/tissue types)
Automated descriptions & expression clusters
- Many expression clusters are associated with infection
- Such data not currently incorporated into automated gene descriptions
- Wen can talk with Ranjana to incorporate (particularly for info-sparse genes)
- Can use GO terms, but need to be careful about how they are applied (what relationships they have) to genes
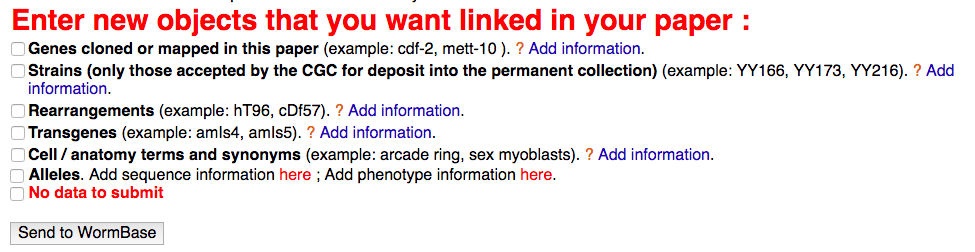
Author First Pass Form
Overall approach
- AFP form would move from being just a flagging pipeline towards a validation and data entry pipeline
- Also move from free text boxes towards autocompletes, controlled vocabularies
- If this is a validation and a portal for data entry forms, should we display everything we can mine?
- Idea is to eventually create data entry forms for as many data types as possible
- Long term goal would include linking to TPC and evidence sentences for validation
- Data types not curated by WB would need to be re-evaluated: continue to mine but share with other curation groups? no longer mine? new WB objects or pages?
- Periodically review the form and add/remove data types and curation forms as needed
- Would be good to ask authors if they had to omit data from the recently published paper and if they want to micropublish
Questions
- Question for WB curators: Are we missing data types?
- Question for WB curators: Are there data types that we don't curate (e.g. domain analysis) but for which we could share mining information with other groups, e.g. UniProt?
- Question for WB curators: Are there data types we should no longer mine?
- Question for Wen: Does it make sense to include an afp flag for expression cluster?
- Question for Karen: What afp tables are populated from the Genetics/G3 pipeline?
- journal first pass forms : http://tazendra.caltech.edu/~azurebrd/cgi-bin/forms/journal/journal_first_pass.cgi?
Details here: http://wiki.wormbase.org/index.php/Genetics_Markup_by_Textpresso_and_First_Pass
Thoughts
- Could we add to citation index score as incentive to provide validation/data?
- How do we make sure we are not turning off/overwhelming users/participants/authors?
- Can gamify the forms; give points?
- Create an app that has user-friendly data submission tools/forms?
December 14, 2017
MOD data in Data Commons
- NIH Data Commons wants DB data in common formats
- We've wanted APIs or web service to allow access all data in standard format
- Good to have one place for all data from all MODs
- This should be a work product of the AGR
- Consider priority: data site, website
- AGR working groups work towards data standards
- GAF (gene association file) could be used now; what can we learn about the GAF: development, usage?
- GAF hasn't changed much in last ~10 years
Contingency plan for Juancarlos' vacation
- If Tazendra crashes/dies, we can perform a hard drive swap at Caltech
Essential genes
- David Fay asked for list of essential C. elegans genes
- David asked for genes whose mutants are inviable as homozygotes, ignoring RNAi
- Could we generate a file for users? Can we make a conservative list with ~no false positives
- What are all the criteria (that are curated) that could be used to decide whether a gene is essential?
- How does any user define "essential"? What are the cutoffs/thresholds?
- Best option may be to provide a table with all relevant phenotype annotation attributes and let user decide based on filters
Expression certainty
- Proteomics experiments have subjective thresholds
- How do we handle cases where one peptide maps to multiple genes?
- How should large scale expression data be incorporated into enrichment analysis?
GitHub tracker for Caltech curation issues
- "Caltech curation" repository - primarily used for creating curation tool set for Caltech team
- "WormBase curation" repository - last used in 2015
- Should there be a single repository for tracking? Yes, use the "wormbase-curation" repository
- Curators will start submitting tickets in the "wormbase-curation" repository
- OA code is in there now
- We should add OA dumper code as well
December 21, 2017
Upload
- Jan 19th citace upload to Hinxton
- Get .ace files to Wen by Tuesday Jan 16th, 10am PST
Wen's outreach slides
- Wen created slides for presenting at San Diego area worm meeting
- Meeting on Jan 12, 2018
- Wen recreated her slides based on feedback from Sternberg lab meeting
- Slides cover step-by-step tutorials for using Wormbase website
- For future outreach meetings/presentations, we can reach out ahead of time to find out what topics would be of greatest interest
- Curators, review Wen's slides and send her feedback
- Can take a look at WormBase YouTube channel for existing videos
- Chris can send WormMine tutorial videos and/or slides to Wen to share with audience
Karen and Daniela going to Bay Area meeting
- January meeting
- WormBase outreach, micropublications
April will start in January
- Will start on transcriptional regulation
- May work on allele phenotypes
Any curation issues (tool bugs etc.)
- While Juancarlos is gone, submit tickets to GitHub
- Raymond will take a look at