Difference between revisions of "Co-op Documentation"
| Line 3: | Line 3: | ||
*'''Refining Interactions widget:''' | *'''Refining Interactions widget:''' | ||
**''Details:'' | **''Details:'' | ||
| − | |||
| − | |||
***Need to scale distance based from current gene based on connectedness(number of interactions) | ***Need to scale distance based from current gene based on connectedness(number of interactions) | ||
****Shouldn't cluster by gene class (problems arise from classes like "let" where genes are necessarily related so grouping is misleading) | ****Shouldn't cluster by gene class (problems arise from classes like "let" where genes are necessarily related so grouping is misleading) | ||
**''Progress:'' | **''Progress:'' | ||
| − | *** | + | **''Related/Example Links:'' |
| − | *** | + | ***[http://dev.wormbase.org:8023/species/c_elegans/gene/WBGene00006763#8--10 unc-26](<del>82</del> 71) |
| + | ***[http://dev.wormbase.org:8023/species/c_elegans/gene/WBGene00000912#8--10 daf-16](<del>433</del> 181) | ||
| + | ***[http://beta.wormbase.org/species/c_elegans/gene/WBGene00000912#8--10 old daf-16 (beta)](1386 w/ duplication) | ||
| + | ***[http://dev.wormbase.org:8023/species/c_elegans/gene/WBGene00000898#8--10 daf-2](600 no nearby) | ||
**''Completed:'' | **''Completed:'' | ||
| + | ***If the number of interactions is too high, only show the minimal table (hide Cytoscape?) and add "Show More"/"Show Neighbouring Interactions"? option | ||
| + | ****Otherwise, show all interactions(nearby interactions included) and Cytoscape plugin | ||
| + | ****NOTE: This may need to be revisited and optimized/cleaned up | ||
| + | ****Looking into BLASTP field in Homology Widget on Protein page that uses similar mechanism | ||
| + | *****remove duplication | ||
| + | *****instead of toggle href load, load diagram with no nearby in toggle | ||
| + | *****create threshold-determining algorithm based on nodes? | ||
***Split interactions into 2 methods, one for those involving current gene and one for nearby interactions | ***Split interactions into 2 methods, one for those involving current gene and one for nearby interactions | ||
| − | ***If local interactions exceeds <del>400</del> 100(after merging duplicates), doesn't load nearby | + | ***If local interactions exceeds <del>400</del> 100(after merging duplicates), doesn't load nearby |
***Combined duplicate edges (duplicate implies matching source node, target node, direction, type, & phenotype) | ***Combined duplicate edges (duplicate implies matching source node, target node, direction, type, & phenotype) | ||
***Changed table: "Details" column with single link to interaction object -> "Citations" column with list of links to duplicate interactions | ***Changed table: "Details" column with single link to interaction object -> "Citations" column with list of links to duplicate interactions | ||
| Line 23: | Line 31: | ||
**''Details:'' | **''Details:'' | ||
***Making corrections to Gene page interface based on curator consultation | ***Making corrections to Gene page interface based on curator consultation | ||
| − | |||
| − | |||
| − | |||
***Phenotype: | ***Phenotype: | ||
****Add evidence information | ****Add evidence information | ||
| Line 37: | Line 42: | ||
***Phenotype: | ***Phenotype: | ||
****Adding evidence info | ****Adding evidence info | ||
| + | **''Related/Example Links:'' | ||
| + | ***[http://dev.wormbase.org:8023/species/c_elegans/gene/WBGene00006763#8--10 unc-26] | ||
**''Completed:'' | **''Completed:'' | ||
***Consultation from curators. Teleconference: 11:30 am EST Wednesday, January 18th, 2012 | ***Consultation from curators. Teleconference: 11:30 am EST Wednesday, January 18th, 2012 | ||
| + | ***Interactions: | ||
| + | ****Added edge-colour legend | ||
| + | ****Hide no_interaction edges unless that option is selected/filtered for by user | ||
***Phenotype: | ***Phenotype: | ||
****Standardize look and feel of tables (convert to data tables) | ****Standardize look and feel of tables (convert to data tables) | ||
Revision as of 19:31, 25 January 2012
Contents
Current Tasks
Tasks In Progress
- Refining Interactions widget:
- Details:
- Need to scale distance based from current gene based on connectedness(number of interactions)
- Shouldn't cluster by gene class (problems arise from classes like "let" where genes are necessarily related so grouping is misleading)
- Need to scale distance based from current gene based on connectedness(number of interactions)
- Progress:
- Related/Example Links:
- unc-26(
8271) - daf-16(
433181) - old daf-16 (beta)(1386 w/ duplication)
- daf-2(600 no nearby)
- unc-26(
- Completed:
- If the number of interactions is too high, only show the minimal table (hide Cytoscape?) and add "Show More"/"Show Neighbouring Interactions"? option
- Otherwise, show all interactions(nearby interactions included) and Cytoscape plugin
- NOTE: This may need to be revisited and optimized/cleaned up
- Looking into BLASTP field in Homology Widget on Protein page that uses similar mechanism
- remove duplication
- instead of toggle href load, load diagram with no nearby in toggle
- create threshold-determining algorithm based on nodes?
- Split interactions into 2 methods, one for those involving current gene and one for nearby interactions
- If local interactions exceeds
400100(after merging duplicates), doesn't load nearby - Combined duplicate edges (duplicate implies matching source node, target node, direction, type, & phenotype)
- Changed table: "Details" column with single link to interaction object -> "Citations" column with list of links to duplicate interactions
- Modified tooltip
- Removed interaction number and link to page (misleading for multiple)
- Added phenotype and number of citations to tooltip
- Added "edge width based on number of citations" feature
- Set 15 as max number of citations; anything higher will be scaled down
- If the number of interactions is too high, only show the minimal table (hide Cytoscape?) and add "Show More"/"Show Neighbouring Interactions"? option
- Details:
- Gene page alterations
- Details:
- Making corrections to Gene page interface based on curator consultation
- Phenotype:
- Add evidence information
- Gene Ontology widget uses similar mechanism
- Check for build_evidence method in Object.pm
- Add evidence information
- Reagents:
- (Replacing altogether by list)
Drop the word "found" for search results - Switch search results to actual list of results(expand if too many)
- See/learn how this was done for the Expression widget
- (Replacing altogether by list)
- Progress:
- Phenotype:
- Adding evidence info
- Phenotype:
- Related/Example Links:
- Completed:
- Consultation from curators. Teleconference: 11:30 am EST Wednesday, January 18th, 2012
- Interactions:
- Added edge-colour legend
- Hide no_interaction edges unless that option is selected/filtered for by user
- Phenotype:
- Standardize look and feel of tables (convert to data tables)
- Done changing Phenotypes, Phenotypes_not_observed, and RNAi tables to use build_data_table
- Modified corresponding API methods to return compatible data for build_data_table
- Standardize look and feel of tables (convert to data tables)
- Details:
- Non-compliant Data Checks
- Details:
- Using fatal_non_compliance check and debug option (enable in wormbase_local.conf) to determine if API methods return invalid data
- E.g. returning ACe objects(usually this should be a string) or empty arrays/hashes(should return undef)
- Usually these can be found when fields do not have data to return but give empty structures instead of just undef
- Using fatal_non_compliance check and debug option (enable in wormbase_local.conf) to determine if API methods return invalid data
- Progress:
- Correcting these as they're encountered
- Completed:
- Methods showing errors on the following pages were corrected:
- Variation, Person, Laboratory, Gene Class, Life Stages, Phenotype, Protein, Strain
- Methods showing errors on the following pages were corrected:
- Details:
- Interface Corrections
- Details:
- Look into rearranging the elements in widgets so that they do not appear strange/distorted when switched from one- to two-column views or using smaller displays
- Will require modifying the templates
- Details:
- Homology Widget
- Needs to be modified; Contains too much information that shouldn't necessarily be grouped this way
- May need to move orthologs to separate widget
- Cytoscape plugin
- Incorporate button/option to open pop-up for larger view (e.g. in case user has small screen or is using 2-column layout)
- Also, add plugin to interaction page (may need to wait until interactions are merged)
Other Tasks
Additional tasks that require further investigation
- Loading/processing large amounts of data
- Details:
- Likely approach will be to change the database used from ACeDB to NoSQL/CouchDB
- Sample Issue
- Progress:
- Ace2Couch Scripts for migrating data
- AceCouch Perl API
- Details:
Getting Started
Members/Contacts
- Abigail Cabunoc - abigail.cabunoc@oicr.on.ca
- Todd Harris - todd@wormbase.org
- Lincoln Stein - lincoln.stein@gmail.com
- Quang Trinh
Meetings
- WormBase OICR Developers Teleconference
- Mondays, 3:00PM
- Phone-in 1-800-747-5150 id: 6738514
- WormBase OICR Developers Teleconference with Lincoln
- Wednesdays, 4:00PM EST
- Lincoln's office
- Agenda and Minutes
- WormBase International Groups Teleconference
- Every other Thursday, 11:30AM
- Lincoln's Office
- Group Meeting
- Fridays, 3:00PM
- HL31 Conference Room/TBA
Tools
The following are some of the tools with which many major aspects of WormBase are developed. You should familiarize yourself with them through documentation and examples.
Perl
There is significant documentation on getting started with Perl. One starting point is PerlMonks.
Catalyst
Catalyst is the web development framework used to develop WormBase. To get started, read and try examples from
- The Definitive Guide to Catalyst (should be available on bookshelf)
- CPAN - About Catalyst
Git
Git is a version control system used for collaboration and backup in the development process. One starting point is the progit tutorial
- WormBase repository located at https://github.com/organizations/WormBase
- Common commands
- status, add, checkout, commit, push, pull, log, fetch, merge
- Usage example:
Given that we have modified two files a.txt and b.txt but do not wish to keep the changes made to b.txt
git status
git checkout b.txt
git add a.txt
git commit -m “Added change1 and change2 to a.txt”
git push
If we have did not have the most recent version, then we will run into an issue when trying to push. In this case we can:
git fetch
git merge
git push
Note: pull is similar to using fetch + merge
Browser (Debugging)
Many browsers provide useful tools to developers that can be used for debugging
Chrome
- Tools->Developer Tools/JavaScript Console
Firefox
- Firefox->Web Developer->Web Console/Error Console
Other
Some other tools that you should be aware of but may not be required to know/interact with include:
- JavaScript
- ACeDB
- MySQL
- Xapian
- GFF
- Cytoscape Web
- Cytoscape Tutorial
- Plugin generally used for pathway analysis
- This plugin is used in the Interactions widget on the Gene page ([WormBase dir]/root/templates/classes/gene/interactions.tt2)
- Installation directory (for updating): [WormBase dir]/root/js/jquery/plugins/
General Concepts
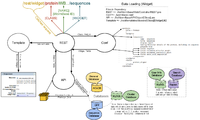
Widget Data Loading
REST Controller:
- Catches internal url (/rest/widget/...)
- Determines the class and widget from the url
- From class and widget, determines which fields are required from the configuration file (wormbase.conf)
- API methods request data from appropriate databases, process/format/package the data, and then return it(/lib/WormBase/API/Object/[Class].pm)
- Sends data to the template to be used in rendering the widget (/root/templates/classes/[Class]/[Widget].tt2)
Other Info
Useful Macros when editing templates can be found in:
- /root/templates/config/main
- /root/templates/shared/page_elements.tt2
Useful files for debugging include:
- /logs/wb-dev-catalyst.log
Other:
- If the port you have been using appears to be busy but the server is not running,
ps -aux|grep -XXXX kill -9 ID
where XXXX is the port number(e.g. 8023) and ID is the process id that is using the port(e.g. 10361) NOTE: Do not kill other users' processes if you are using a shared dev machine (e.g. wb-dev)
- To dump data in API methods include Data::Dumper
use Data::Dumper ... warn(Dumper(\@data));
- Sometimes firewall blocks certain ports, so you may be unable to connect to your dev server on the wb-dev machine through the browser
ssh -L 8080:localhost:XXXX wb-dev.oicr.on.ca ./wormbase_server.pl -p XXXX -d -r
Then go to localhost:8080 in browser